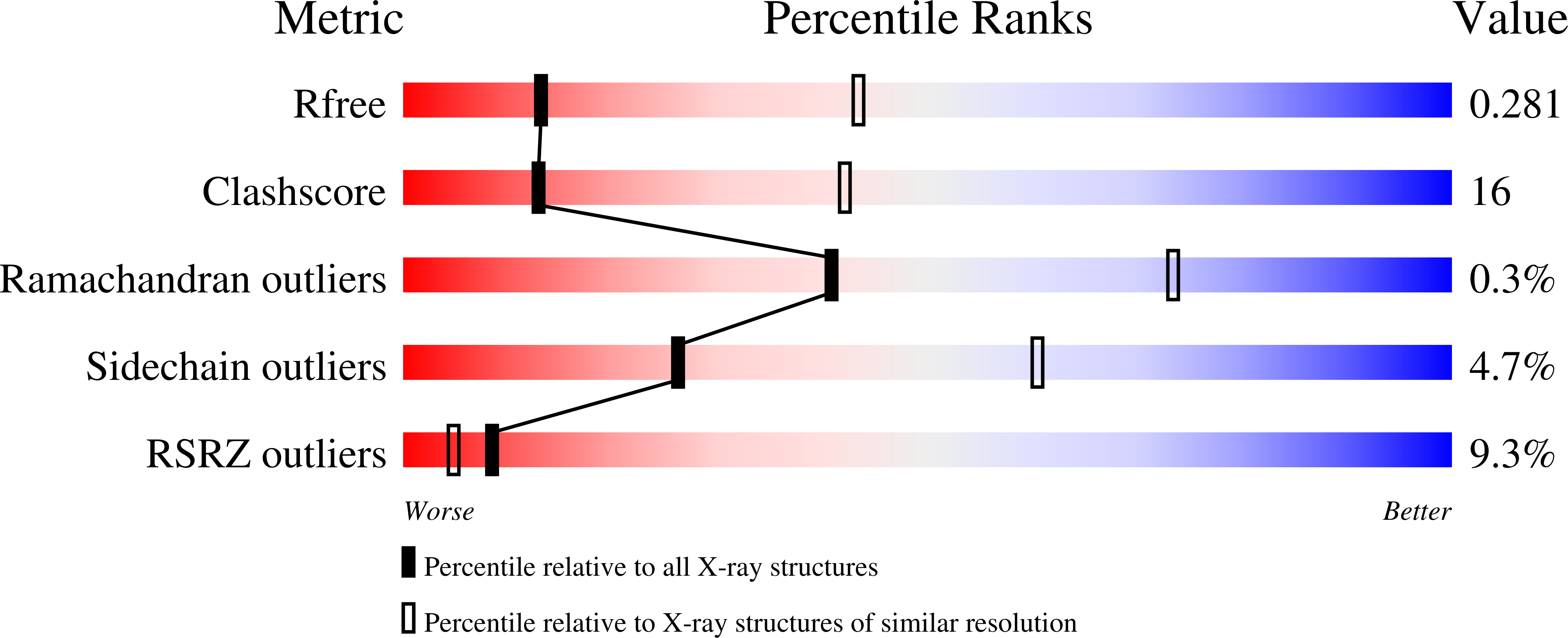
Binding of a physiological substrate causes large-scale conformational reorganization in cytochrome P450 51.
Hargrove, T.Y., Wawrzak, Z., Fisher, P.M., Child, S.A., Nes, W.D., Guengerich, F.P., Waterman, M.R., Lepesheva, G.I.(2018) J Biol Chem 293: 19344-19353
- PubMed: 30327430
- DOI: https://doi.org/10.1074/jbc.RA118.005850
- Primary Citation of Related Structures:
6FMO - PubMed Abstract:
Sterol 14α-demethylases (CYP51s) are phylogenetically the most conserved cytochromes P450, and their three-step reaction is crucial for biosynthesis of sterols and serves as a leading target for clinical and agricultural antifungal agents. The structures of several (bacterial, protozoan, fungal, and human) CYP51 orthologs, in both the ligand-free and inhibitor-bound forms, have been determined and have revealed striking similarity at the secondary and tertiary structural levels, despite having low sequence identity. Moreover, in contrast to many of the substrate-promiscuous, drug-metabolizing P450s, CYP51 structures do not display substantial rearrangements in their backbones upon binding of various inhibitory ligands, essentially representing a snapshot of the ligand-free sterol 14α-demethylase. Here, using the obtusifoliol-bound I105F variant of Trypanosoma cruzi CYP51, we report that formation of the catalytically competent complex with the physiological substrate triggers a large-scale conformational switch, dramatically reshaping the enzyme active site (3.5-6.0 Å movements in the FG arm, HI arm, and helix C) in the direction of catalysis. Notably, our X-ray structural analyses revealed that the substrate channel closes, the proton delivery route opens, and the topology and electrostatic potential of the proximal surface reorganize to favor interaction with the electron-donating flavoprotein partner, NADPH-cytochrome P450 reductase. Site-directed mutagenesis of the amino acid residues involved in these events revealed a key role of active-site salt bridges in contributing to the structural dynamics that accompanies CYP51 function. Comparative analysis of apo-CYP51 and its sterol-bound complex provided key conceptual insights into the molecular mechanisms of CYP51 catalysis, functional conservation, lineage-specific substrate complementarity, and druggability differences.
Organizational Affiliation:
From the Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, Tennessee 37232.