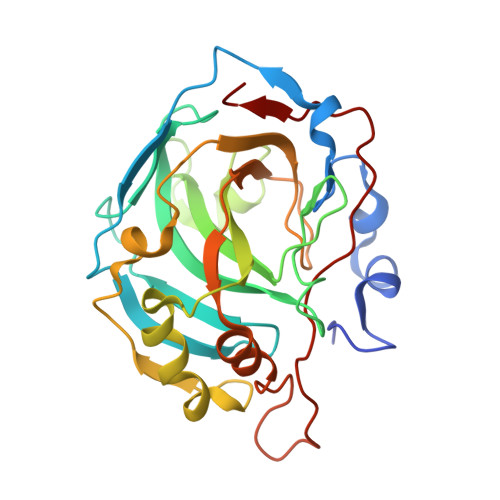
Design of two-tail compounds with rotationally fixed benzenesulfonamide ring as inhibitors of carbonic anhydrases.
Zaksauskas, A., Capkauskaite, E., Jezepcikas, L., Linkuviene, V., Kisonaite, M., Smirnov, A., Manakova, E., Grazulis, S., Matulis, D.(2018) Eur J Med Chem 156: 61-78
- PubMed: 30006175
- DOI: https://doi.org/10.1016/j.ejmech.2018.06.059
- Primary Citation of Related Structures:
6G5L, 6G5U, 6G6T, 6G7A - PubMed Abstract:
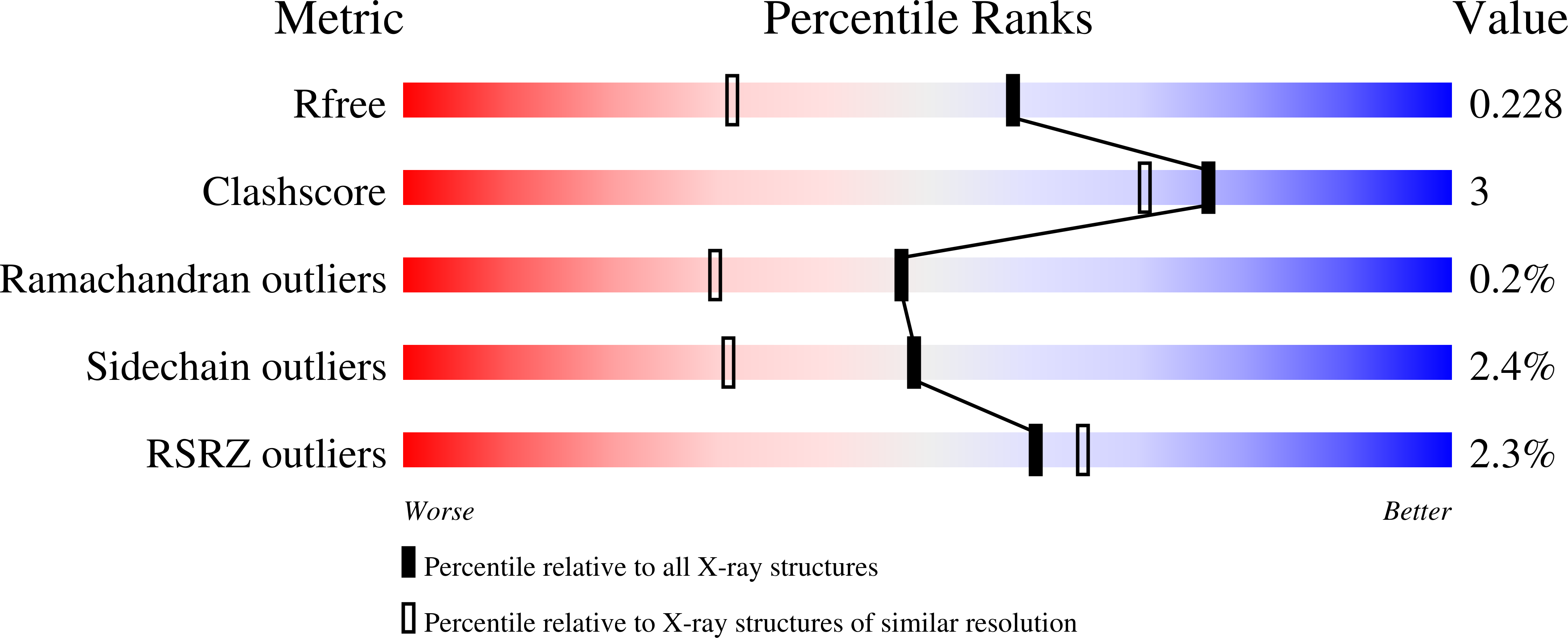
Rational design of compounds that would bind specific pockets of the target proteins is a difficult task in drug design. The 12 isoforms of catalytically active human carbonic anhydrases (CAs) have highly similar active sites that make it difficult to design inhibitors selective for one or several CA isoforms. A series of CA inhibitors based on 2-chloro/bromo-benzenesulfonamide that is largely fixed in the CA active site together with one or two tails yielded compounds that were synthesized and evaluated as inhibitors of CA isoforms. Introduction of a second tail had significant influence on the binding affinity and two-tailed compounds in most cases provided high affinity and selectivity for CA IX and CA XIV. The contacts between several compounds and CA amino acids were determined by X-ray crystallography. Together with the intrinsic enthalpy and entropy of binding they provided the structure-thermodynamics correlations for this series of compounds with the insight how to rationally build compounds with desired CA isoform as a target.
Organizational Affiliation:
Department of Biothermodynamics and Drug Design, Institute of Biotechnology, Vilnius University, Saulėtekio al. 7, Vilnius LT-10257, Lithuania.