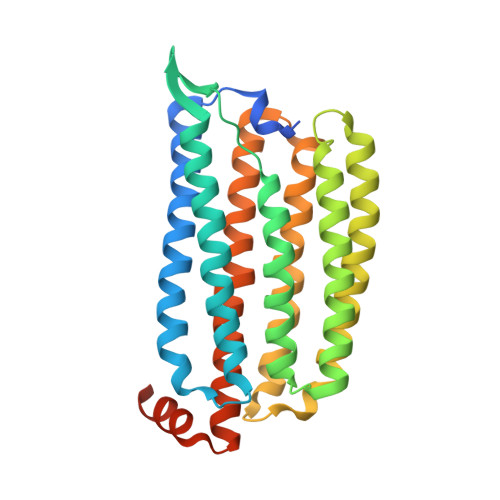
Pumping mechanism of NM-R3, a light-driven bacterial chloride importer in the rhodopsin family.
Yun, J.H., Ohki, M., Park, J.H., Ishimoto, N., Sato-Tomita, A., Lee, W., Jin, Z., Tame, J.R.H., Shibayama, N., Park, S.Y., Lee, W.(2020) Sci Adv 6: eaay2042-eaay2042
- PubMed: 32083178
- DOI: https://doi.org/10.1126/sciadv.aay2042
- Primary Citation of Related Structures:
6JY6, 6JY7, 6JY8, 6JY9, 6JYA, 6JYB, 6JYC, 6JYD, 6JYE, 6JYF - PubMed Abstract:
A newly identified microbial rhodopsin, NM-R3, from the marine flavobacterium Nonlabens marinus , was recently shown to drive chloride ion uptake, extending our understanding of the diversity of mechanisms for biological energy conversion. To clarify the mechanism underlying its function, we characterized the crystal structures of NM-R3 in both the dark state and early intermediate photoexcited states produced by laser pulses of different intensities and temperatures. The displacement of chloride ions at five different locations in the model reflected the detailed anion-conduction pathway, and the activity-related key residues-Cys 105 , Ser 60 , Gln 224 , and Phe 90 -were identified by mutation assays and spectroscopy. Comparisons with other proteins, including a closely related outward sodium ion pump, revealed key motifs and provided structural insights into light-driven ion transport across membranes by the NQ subfamily of rhodopsins. Unexpectedly, the response of the retinal in NM-R3 to photostimulation appears to be substantially different from that seen in bacteriorhodopsin.
Organizational Affiliation:
Department of Biochemistry, College of Life Science and Biotechnology, Yonsei University, Seoul 03722, South Korea.