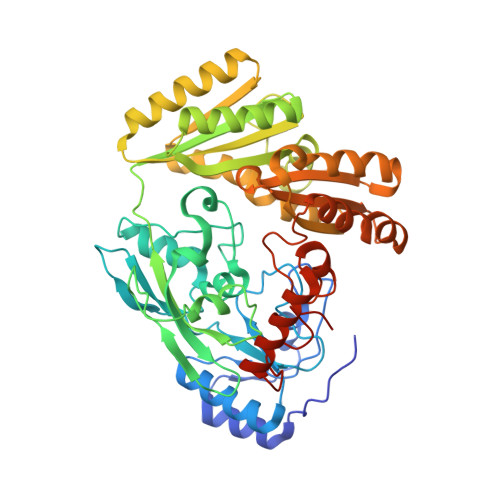
Structure, substrate specificity, and catalytic mechanism of human D-2-HGDH and insights into pathogenicity of disease-associated mutations.
Yang, J., Zhu, H., Zhang, T., Ding, J.(2021) Cell Discov 7: 3-3
- PubMed: 33431826
- DOI: https://doi.org/10.1038/s41421-020-00227-0
- Primary Citation of Related Structures:
6LPN, 6LPP, 6LPQ, 6LPT, 6LPU, 6LPX - PubMed Abstract:
D-2-hydroxyglutarate dehydrogenase (D-2-HGDH) catalyzes the oxidation of D-2-hydroxyglutarate (D-2-HG) into 2-oxoglutarate, and genetic D-2-HGDH deficiency leads to abnormal accumulation of D-2-HG which causes type I D-2-hydroxyglutaric aciduria and is associated with diffuse large B-cell lymphoma. This work reports the crystal structures of human D-2-HGDH in apo form and in complexes with D-2-HG, D-malate, D-lactate, L-2-HG, and 2-oxoglutarate, respectively. D-2-HGDH comprises a FAD-binding domain, a substrate-binding domain, and a small C-terminal domain. The active site is located at the interface of the FAD-binding domain and the substrate-binding domain. The functional roles of the key residues involved in the substrate binding and catalytic reaction and the mutations identified in D-2-HGDH-deficient diseases are analyzed by biochemical studies. The structural and biochemical data together reveal the molecular mechanism of the substrate specificity and catalytic reaction of D-2-HGDH and provide insights into the pathogenicity of the disease-associated mutations.
Organizational Affiliation:
State Key Laboratory of Molecular Biology, Shanghai Institute of Biochemistry and Cell Biology, Center for Excellence in Molecular Cell Science, University of Chinese Academy of Sciences, Chinese Academy of Sciences, 320 Yue-Yang Road, Shanghai, 200031, China.