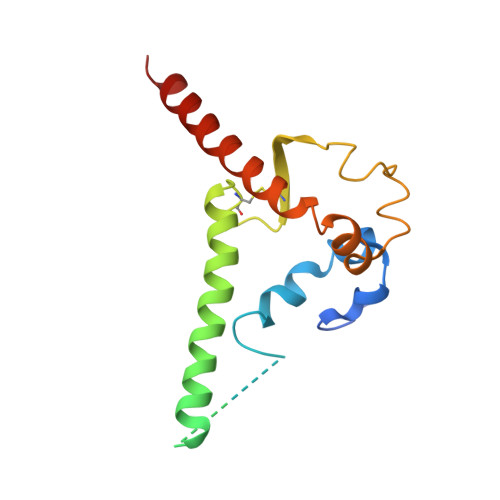
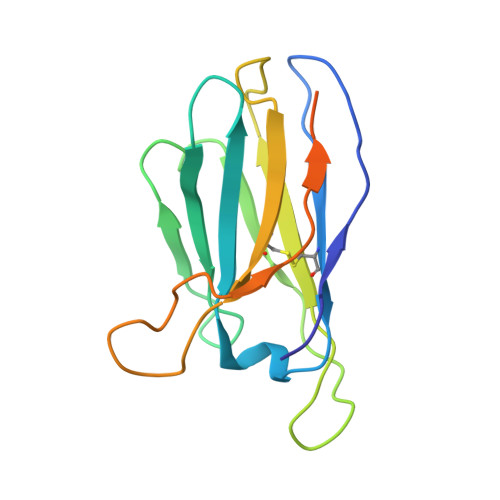
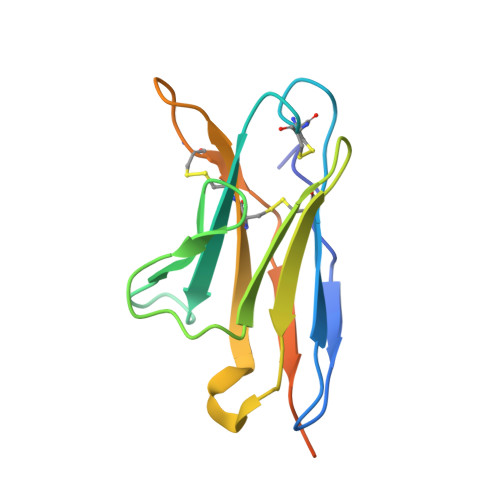
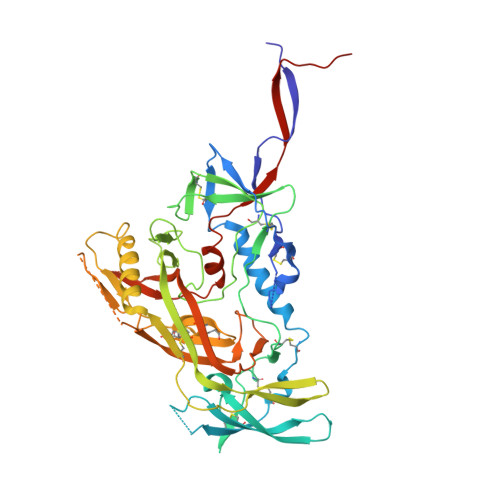
Improvement of antibody functionality by structure-guided paratope engraftment.
Liu, Q., Lai, Y.T., Zhang, P., Louder, M.K., Pegu, A., Rawi, R., Asokan, M., Chen, X., Shen, C.H., Chuang, G.Y., Yang, E.S., Miao, H., Wang, Y., Fauci, A.S., Kwong, P.D., Mascola, J.R., Lusso, P.(2019) Nat Commun 10: 721-721
- PubMed: 30760721
- DOI: https://doi.org/10.1038/s41467-019-08658-4
- Primary Citation of Related Structures:
6NM6, 6NNF, 6NNJ - PubMed Abstract:
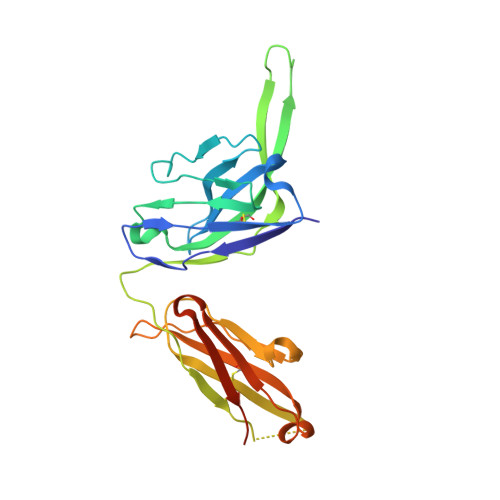
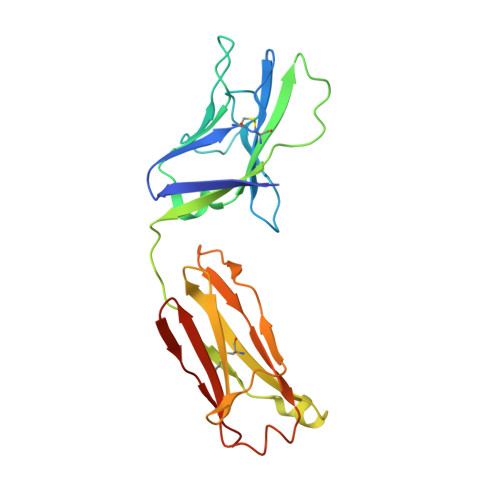
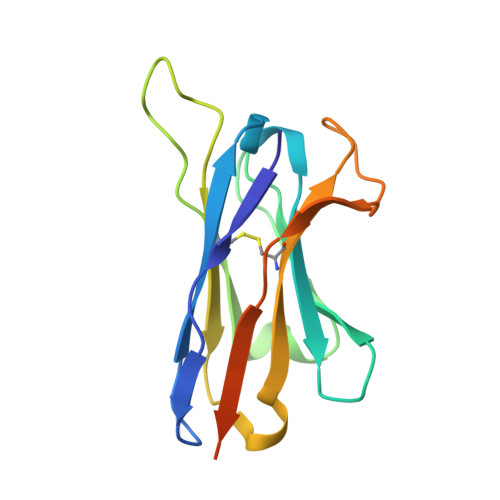
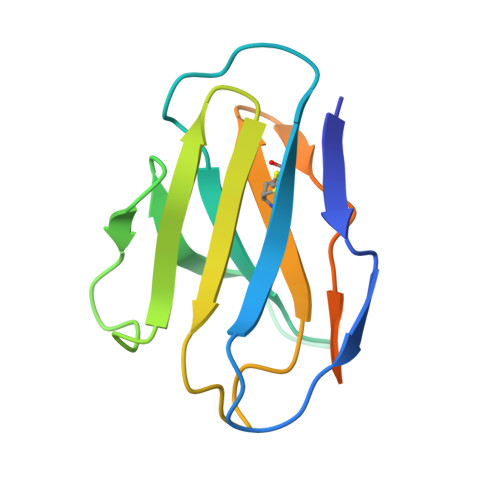
Broadly neutralizing antibodies (bNAbs) represent a promising alternative to antiretroviral drugs for HIV-1 prevention and treatment. Selected antibodies to the CD4-binding site bolster envelope trimer binding via quaternary contacts. Here, we rationally engraft a new paratope, i.e., the extended heavy-chain framework region 3 (FR3) loop of VRC03, which mediates quaternary interaction, onto several potent bNAbs, enabling them to reach an adjacent gp120 protomer. The interactive quaternary surface is delineated by solving the crystal structure of two FR3 loop-chimeric antibodies. Chimerization enhances the neutralizing activity of several potent bNAbs against a majority of global HIV-1 strains. Compared to unmodified antibodies, chimeric antibodies display lower autoreactivity and prolonged in vivo half-life in huFcRn mice and rhesus macaques. Thus, paratope engraftment may be used to expand the epitope repertory of natural antibodies, improving their functionality for disease prevention and treatment.
Organizational Affiliation:
Laboratory of Immunoregulation, National Institute of Allergy and Infectious Diseases, NIH, Bethesda, MD, 20892, USA.