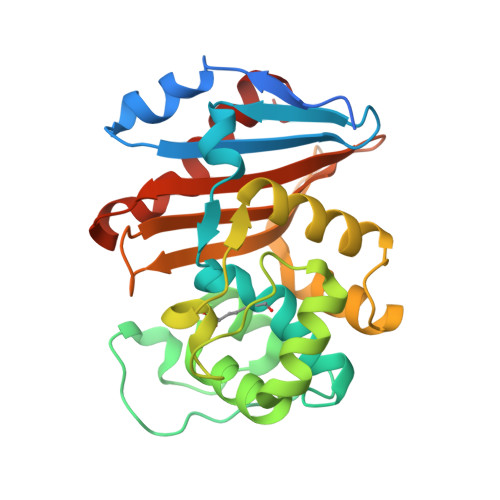
Structural Basis for Substrate Specificity and Carbapenemase Activity of OXA-48 Class D beta-Lactamase.
Akhtar, A., Pemberton, O.A., Chen, Y.(2020) ACS Infect Dis 6: 261-271
- PubMed: 31872762
- DOI: https://doi.org/10.1021/acsinfecdis.9b00304
- Primary Citation of Related Structures:
6PK0, 6PQI, 6PSG, 6PT1, 6PT5, 6PTU - PubMed Abstract:
Carbapenem-hydrolyzing class D β-lactamases (CHDLs) are a diverse family of enzymes that are rapidly becoming the predominant cause of bacterial resistance against β-lactam antibiotics in many regions of the world. OXA-48, an atypical member of CHDLs, is one of the most frequently observed in the clinic and exhibits a unique substrate profile. We applied X-ray crystallography to OXA-48 complexes with multiple β-lactam antibiotics to elucidate this enzyme's carbapenemase activity and its preference of imipenem over meropenem and other substrates such as cefotaxime. In particular, we obtained acyl-enzyme complexes of OXA-48 with imipenem, meropenem, faropenem, cefotaxime, and cefoxitin, and a product complex with imipenem. Importantly, the product complex captures a key reaction milestone with the newly generated carboxylate group still in the oxyanion hole, and represents the first such complex with a wild-type serine β-lactamase. A potential hydrogen bond is observed between the two carboxylate groups from the product and the carbamylated Lys73, representing the stage immediately after the breakage of the acyl-enzyme bond where the product carboxylate would be neutral. The placement of the product carboxylate also illustrates the approximate transient location of the deacylation water that has long eluded structural characterization in class D β-lactamases. Additionally, comparing the product complex with the acyl-enzyme intermediates provides new insights into the various mechanisms by which specific side chain groups hinder the access of the deacylation water to the acyl-enzyme linkage, especially in meropenem. Taken together, these data offer valuable information on the substrate specificity of OXA-48 and the catalytic mechanism of CHDLs.
Organizational Affiliation:
Department of Molecular Medicine , University of South Florida Morsani College of Medicine , 12901 Bruce B. Downs Boulevard, MDC 3522 , Tampa , Florida 33612 , United States.