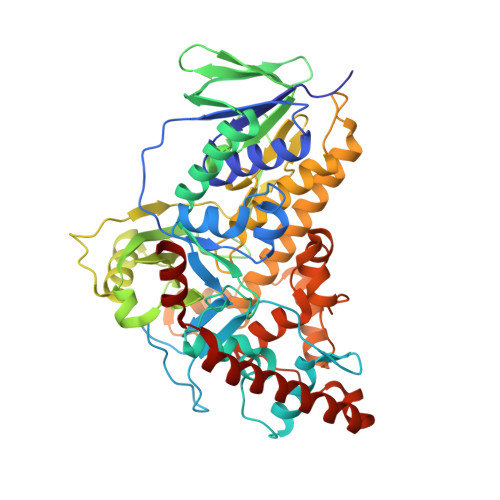
Binding of FAD and tryptophan to the tryptophan 6-halogenase Thal is negatively coupled.
Moritzer, A.C., Niemann, H.H.(2019) Protein Sci 28: 2112-2118
- PubMed: 31589794
- DOI: https://doi.org/10.1002/pro.3739
- Primary Citation of Related Structures:
6SLS, 6SLT - PubMed Abstract:
Flavin-dependent halogenases require reduced flavin adenine dinucleotide (FADH 2 ), O 2 , and halide salts to halogenate their substrates. We describe the crystal structures of the tryptophan 6-halogenase Thal in complex with FAD or with both tryptophan and FAD. If tryptophan and FAD were soaked simultaneously, both ligands showed impaired binding and in some cases only the adenosine monophosphate or the adenosine moiety of FAD was resolved, suggesting that tryptophan binding increases the mobility mainly of the flavin mononucleotide moiety. This confirms a negative cooperativity between the binding of substrate and cofactor that was previously described for other tryptophan halogenases. Binding of substrate to tryptophan halogenases reduces the affinity for the oxidized cofactor FAD presumably to facilitate the regeneration of FADH 2 by flavin reductases.
Organizational Affiliation:
Department of Chemistry, Bielefeld University, Bielefeld, Germany.