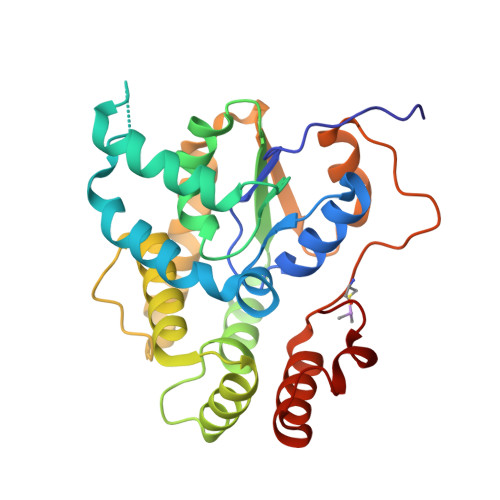
Molecular basis for hycanthone drug action in schistosome parasites.
Guzman, M., Rugel, A., Tarpley, R.S., Cao, X., McHardy, S.F., LoVerde, P.T., Taylor, A.B.(2020) Mol Biochem Parasitol 236: 111257-111257
- PubMed: 32027942
- DOI: https://doi.org/10.1016/j.molbiopara.2020.111257
- Primary Citation of Related Structures:
6UUX, 6UUY - PubMed Abstract:
Hycanthone (HYC) is a retired drug formerly used to treat schistosomiasis caused by infection from Schistosoma mansoni and S. haematobium. Resistance to HYC was first observed in S. mansoni laboratory strains and in patients in the 1970s and the use of this drug was subsequently discontinued with the substitution of praziquantel (PZQ) as the single antischistosomal drug in the worldwide formulary. In endemic regions, multiple organizations have partnered with the World Health Organization to deliver PZQ for morbidity control and prevention. While the monotherapy reduces the disease burden, additional drugs are needed to use in combination with PZQ to stay ahead of potential drug resistance. HYC will not be reintroduced into the schistosomiasis drug formulary as a combination drug because it was shown to have adverse properties including mutagenic, teratogenic and carcinogenic activities. Oxamniquine (OXA) was used to treat S. mansoni infection in Brazil during the brief period of HYC use, until the 1990s. Its antischistosomal efficacy has been shown to work through the same mechanism as HYC and it does not possess the undesirable properties linked to HYC. OXA demonstrates cross-resistance in Schistosoma strains with HYC resistance and both are prodrugs requiring metabolic activation in the worm to toxic sulfated forms. The target activating enzyme has been identified as a sulfotransferase enzyme and is currently used as the basis for a structure-guided drug design program. Here, we characterize the sulfotransferases from S. mansoni and S. haematobium in complexes with HYC to compare and contrast with OXA-bound sulfotransferase crystal structures. Although HYC is discontinued for antischistosomal treatment, it can serve as a resource for design of derivative compounds without contraindication.
Organizational Affiliation:
Department of Pathology and Laboratory Medicine, University of Texas Health Science Center at San Antonio, San Antonio, TX 78229, United States; Department of Microbiology, Immunology & Molecular Genetics, University of Texas Health Science Center at San Antonio, San Antonio, TX 78229, United States; Department of Biochemistry & Structural Biology, University of Texas Health Science Center at San Antonio, San Antonio, TX 78229, United States.