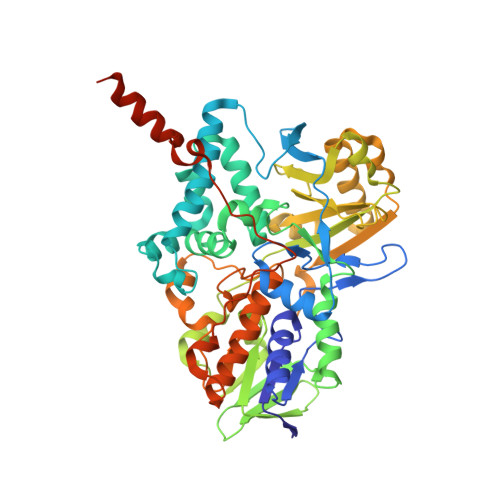
Diphenylene Iodonium Is a Noncovalent MAO Inhibitor: A Biochemical and Structural Analysis.
Iacovino, L.G., Reis, J., Mai, A., Binda, C., Mattevi, A.(2020) ChemMedChem 15: 1394-1397
- PubMed: 32459875
- DOI: https://doi.org/10.1002/cmdc.202000264
- Primary Citation of Related Structures:
6YT2 - PubMed Abstract:
Diphenylene iodonium (DPI) is known for its inhibitory activities against many flavin- and heme-dependent enzymes, and is often used as an NADPH oxidase inhibitor. We probed the efficacy of DPI on two well-known drug targets, the human monoamine oxidases MAO A and B. UV-visible spectrophotometry and steady-state kinetics experiments demonstrate that DPI acts as a competitive and reversible MAO inhibitor with K i values of 1.7 and 0.3 μM for MAO A and MAO B, respectively. Elucidation of the crystal structure of human MAO B bound to the inhibitor revealed that DPI binds deeply in the active-site cavity to establish multiple hydrophobic interactions with the surrounding side chains and the flavin. These data prove that DPI is a genuine MAO inhibitor and that the inhibition mechanism does not involve a reaction with the reduced flavin. This binding and inhibitory activity against the MAOs, two major reactive oxygen species (ROS)-producing enzymes, will have to be carefully considered when interpreting experiments that rely on DPI for target validation and chemical biology studies on ROS functions.
Organizational Affiliation:
Department of Biology and Biotechnology "Lazzaro Spallanzani", University of Pavia, Via Ferrata 9, 27100, Pavia, Italy.