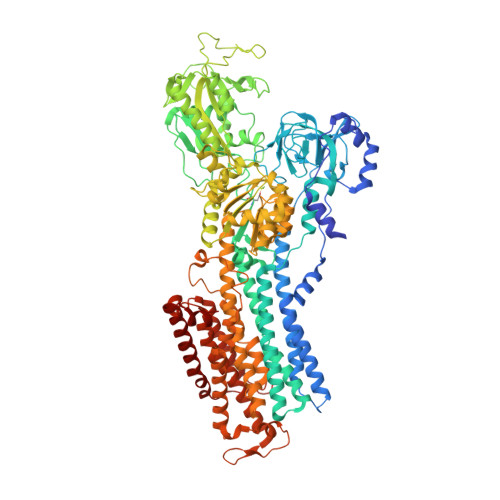
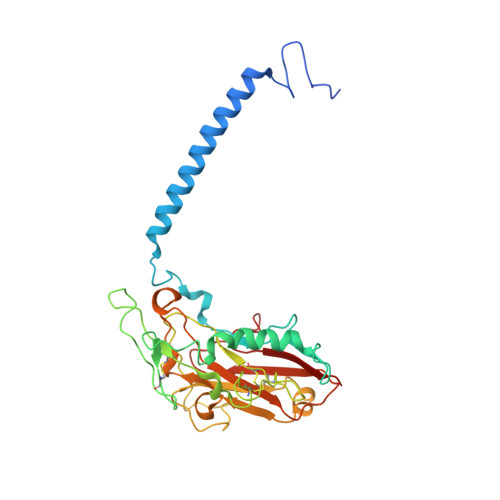
Binding of cardiotonic steroids to Na + ,K + -ATPase in the E2P state.
Kanai, R., Cornelius, F., Ogawa, H., Motoyama, K., Vilsen, B., Toyoshima, C.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33318128
- DOI: https://doi.org/10.1073/pnas.2020438118
- Primary Citation of Related Structures:
7D91, 7D92, 7D93, 7D94, 7DDF, 7DDH, 7DDI, 7DDK, 7DDL - PubMed Abstract:
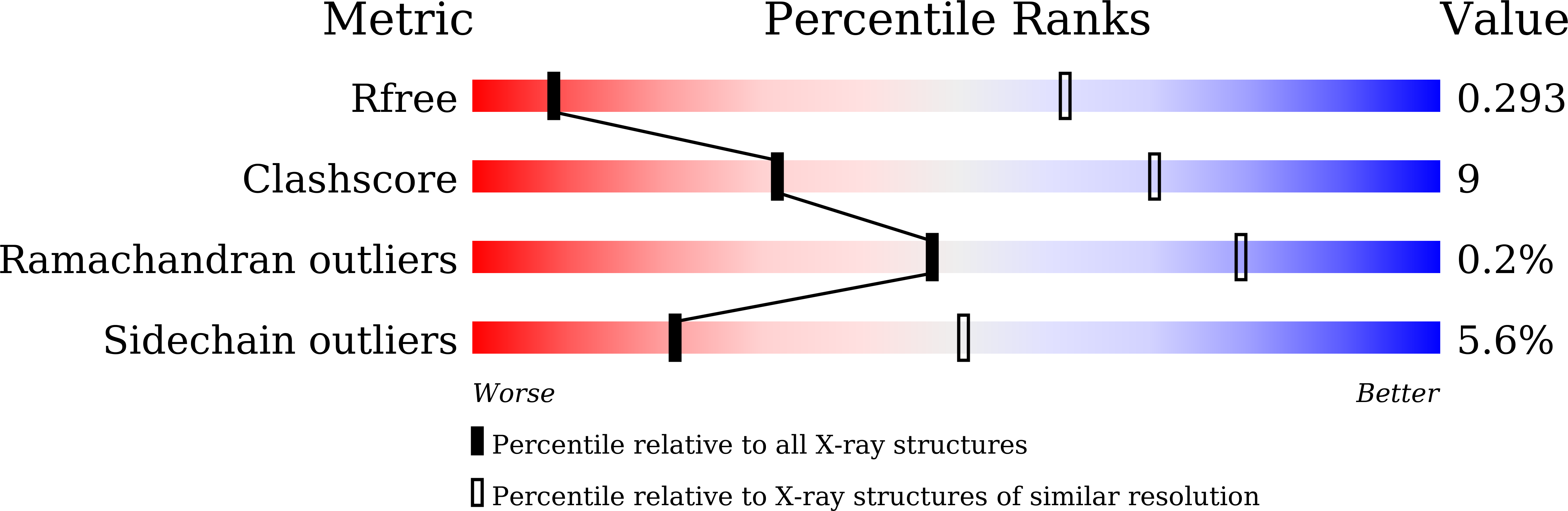
The sodium pump (Na + , K + -ATPase, NKA) is vital for animal cells, as it actively maintains Na + and K + electrochemical gradients across the cell membrane. It is a target of cardiotonic steroids (CTSs) such as ouabain and digoxin. As CTSs are almost unique strong inhibitors specific to NKA, a wide range of derivatives has been developed for potential therapeutic use. Several crystal structures have been published for NKA-CTS complexes, but they fail to explain the largely different inhibitory properties of the various CTSs. For instance, although CTSs are thought to inhibit ATPase activity by binding to NKA in the E2P state, we do not know if large conformational changes accompany binding, as no crystal structure is available for the E2P state free of CTS. Here, we describe crystal structures of the BeF 3 - complex of NKA representing the E2P ground state and then eight crystal structures of seven CTSs, including rostafuroxin and istaroxime, two new members under clinical trials, in complex with NKA in the E2P state. The conformations of NKA are virtually identical in all complexes with and without CTSs, showing that CTSs bind to a preformed cavity in NKA. By comparing the inhibitory potency of the CTSs measured under four different conditions, we elucidate how different structural features of the CTSs result in different inhibitory properties. The crystal structures also explain K + -antagonism and suggest a route to isoform specific CTSs.
Organizational Affiliation:
Institute for Quantitative Biosciences, The University of Tokyo, 113-0032 Tokyo, Japan.