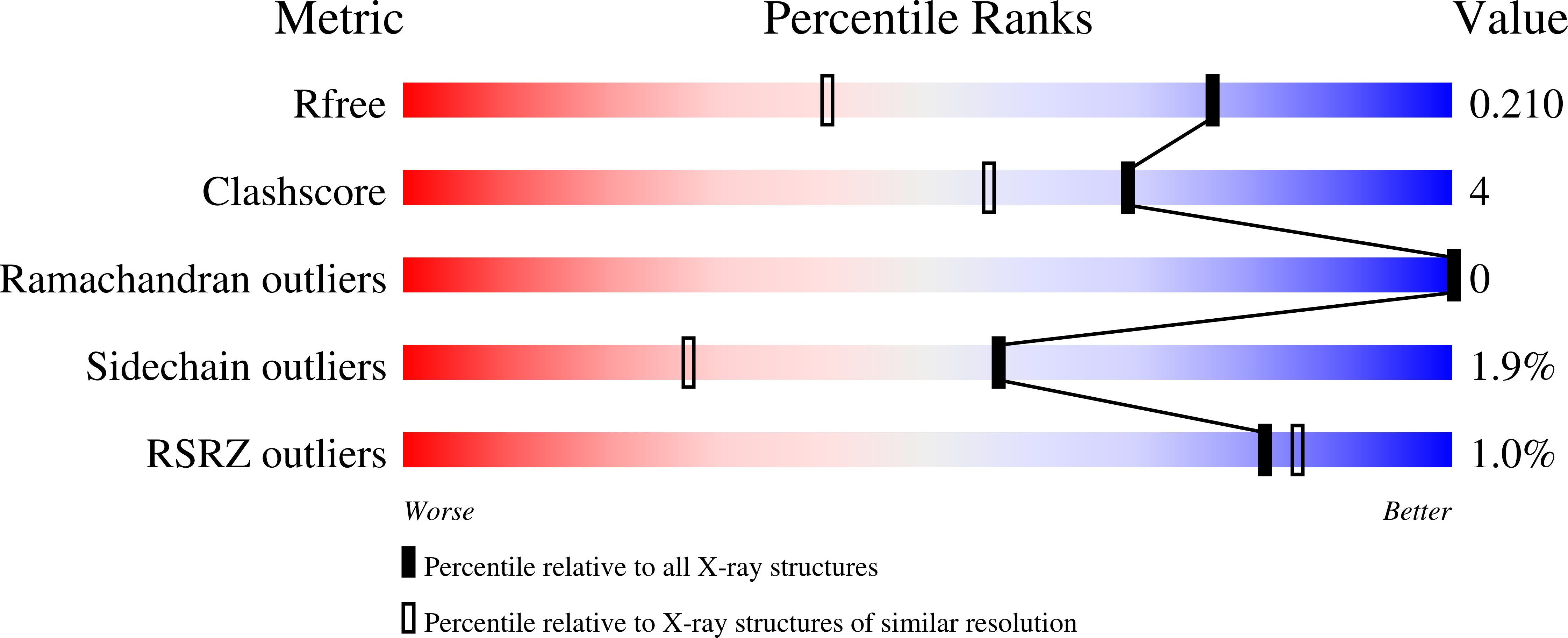
Diverse MarR bacterial regulators of auxin catabolism in the plant microbiome.
Conway, J.M., Walton, W.G., Salas-Gonzalez, I., Law, T.F., Lindberg, C.A., Crook, L.E., Kosina, S.M., Fitzpatrick, C.R., Lietzan, A.D., Northen, T.R., Jones, C.D., Finkel, O.M., Redinbo, M.R., Dangl, J.L.(2022) Nat Microbiol 7: 1817-1833
- PubMed: 36266335
- DOI: https://doi.org/10.1038/s41564-022-01244-3
- Primary Citation of Related Structures:
7KFO, 7KFQ, 7KFS, 7KH3, 7KIG, 7KJL, 7KJQ, 7KK0, 7KKC, 7KKI, 7KRH, 7KUA, 7KYM, 7L19, 7L1I - PubMed Abstract:
Chemical signalling in the plant microbiome can have drastic effects on microbial community structure, and on host growth and development. Previously, we demonstrated that the auxin metabolic signal interference performed by the bacterial genus Variovorax via an auxin degradation locus was essential for maintaining stereotypic root development in an ecologically relevant bacterial synthetic community. Here, we dissect the Variovorax auxin degradation locus to define the genes iadDE as necessary and sufficient for indole-3-acetic acid (IAA) degradation and signal interference. We determine the crystal structures and binding properties of the operon's MarR-family repressor with IAA and other auxins. Auxin degradation operons were identified across the bacterial tree of life and we define two distinct types on the basis of gene content and metabolic products: iac-like and iad-like. The structures of MarRs from representatives of each auxin degradation operon type establish that each has distinct IAA-binding pockets. Comparison of representative IAA-degrading strains from diverse bacterial genera colonizing Arabidopsis plants show that while all degrade IAA, only strains containing iad-like auxin-degrading operons interfere with auxin signalling in a complex synthetic community context. This suggests that iad-like operon-containing bacterial strains, including Variovorax species, play a key ecological role in modulating auxins in the plant microbiome.
Organizational Affiliation:
Department of Biology, University of North Carolina at Chapel Hill, Chapel Hill, NC, USA.