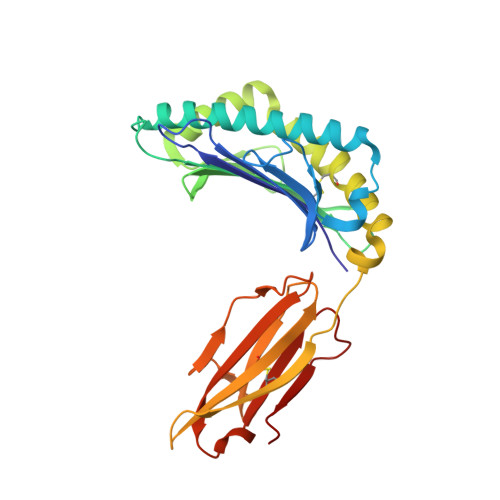
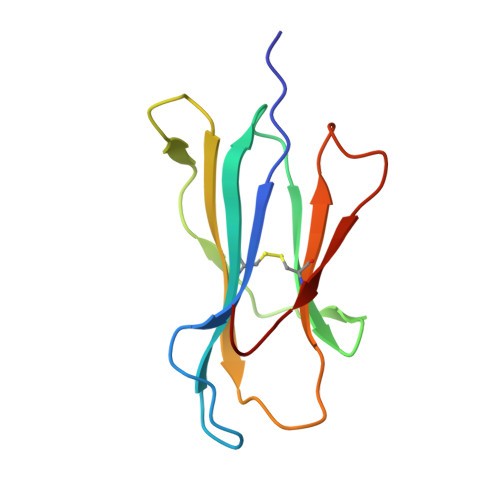
Targeting of multiple tumor-associated antigens by individual T cell receptors during successful cancer immunotherapy.
Dolton, G., Rius, C., Wall, A., Szomolay, B., Bianchi, V., Galloway, S.A.E., Hasan, M.S., Morin, T., Caillaud, M.E., Thomas, H.L., Theaker, S., Tan, L.R., Fuller, A., Topley, K., Legut, M., Attaf, M., Hopkins, J.R., Behiry, E., Zabkiewicz, J., Alvares, C., Lloyd, A., Rogers, A., Henley, P., Fegan, C., Ottmann, O., Man, S., Crowther, M.D., Donia, M., Svane, I.M., Cole, D.K., Brown, P.E., Rizkallah, P., Sewell, A.K.(2023) Cell 186: 3333-3349.e27
- PubMed: 37490916
- DOI: https://doi.org/10.1016/j.cell.2023.06.020
- Primary Citation of Related Structures:
7Q98, 7Q99, 7Q9A, 7Q9B, 7ZUC - PubMed Abstract:
The T cells of the immune system can target tumors and clear solid cancers following tumor-infiltrating lymphocyte (TIL) therapy. We used combinatorial peptide libraries and a proteomic database to reveal the antigen specificities of persistent cancer-specific T cell receptors (TCRs) following successful TIL therapy for stage IV malignant melanoma. Remarkably, individual TCRs could target multiple different tumor types via the HLA A ∗ 02:01-restricted epitopes EAAGIGILTV, LLLGIGILVL, and NLSALGIFST from Melan A, BST2, and IMP2, respectively. Atomic structures of a TCR bound to all three antigens revealed the importance of the shared x-x-x-A/G-I/L-G-I-x-x-x recognition motif. Multi-epitope targeting allows individual T cells to attack cancer in several ways simultaneously. Such "multipronged" T cells exhibited superior recognition of cancer cells compared with conventional T cell recognition of individual epitopes, making them attractive candidates for the development of future immunotherapies.
Organizational Affiliation:
Division of Infection and Immunity, Cardiff University School of Medicine, Cardiff, Wales CF14 4XN, UK.