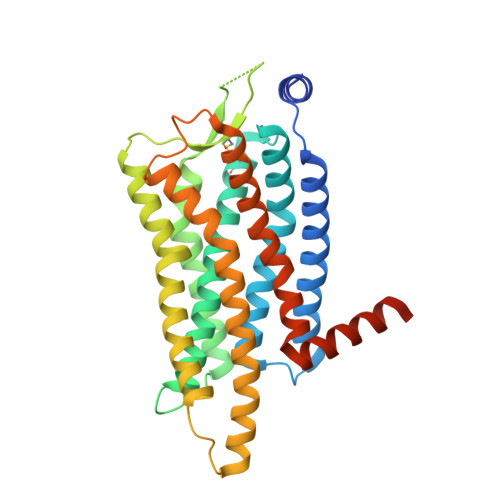
Molecular basis for anti-insomnia drug design from structure of lemborexant-bound orexin 2 receptor.
Asada, H., Im, D., Hotta, Y., Yasuda, S., Murata, T., Suno, R., Iwata, S.(2022) Structure 30: 1582-1589.e4
- PubMed: 36417909
- DOI: https://doi.org/10.1016/j.str.2022.11.001
- Primary Citation of Related Structures:
7XRR - PubMed Abstract:
Orexin receptors are a family of G protein-coupled receptors that consist of two subtypes: orexin-1 receptors (OX1Rs) and OX2Rs. They are expressed throughout the central nervous system and are involved in regulating the sleep-wake cycle. The development of antagonists to orexin receptors has become important in drug discovery because modulation of these receptors can lead to novel treatments for diseases related to the regulation of sleep and wakefulness, such as insomnia. In this study, we determined that the structure of OX2R bound to lemborexant, a dual orexin receptor antagonist (DORA), at 2.89 Å resolution. Comparisons of kinetic and dynamic properties of DORAs based on structures and simulations suggest that the enthalpy of molecular binding to receptors and the entropy derived from intramolecular structure can be separately controlled. These results complement existing structural information and allow us to discuss the usefulness of pharmacophore models and target selectivity to OXRs.
Organizational Affiliation:
Department of Cell Biology, Graduate School of Medicine, Kyoto University, Kyoto 606-8501, Japan. Electronic address: [email protected].