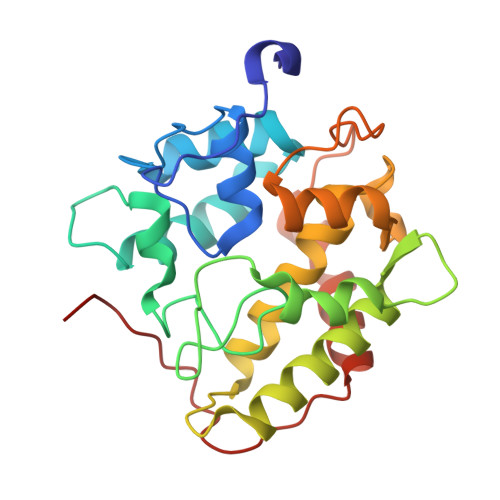
Structural Characterization of Two Short Unspecific Peroxygenases: Two Different Dimeric Arrangements.
Linde, D., Santillana, E., Fernandez-Fueyo, E., Gonzalez-Benjumea, A., Carro, J., Gutierrez, A., Martinez, A.T., Romero, A.(2022) Antioxidants (Basel) 11
- PubMed: 35624755
- DOI: https://doi.org/10.3390/antiox11050891
- Primary Citation of Related Structures:
7ZBP, 7ZCL - PubMed Abstract:
Unspecific peroxygenases (UPOs) are extracellular fungal enzymes of biotechnological interest as self-sufficient (and more stable) counterparts of cytochrome P450 monooxygenases, the latter being present in most living cells. Expression hosts and structural information are crucial for exploiting UPO diversity (over eight thousand UPO-type genes were identified in sequenced genomes) in target reactions of industrial interest. However, while many thousands of entries in the Protein Data Bank include molecular coordinates of P450 enzymes, only 19 entries correspond to UPO enzymes, and UPO structures from only two species ( Agrocybe aegerita and Hypoxylon sp.) have been published to date. In the present study, two UPOs from the basidiomycete Marasmius rotula (r Mro UPO) and the ascomycete Collariella virescens (r Cvi UPO) were crystallized after sequence optimization and Escherichia coli expression as active soluble enzymes. Crystals of r Mro UPO and r Cvi UPO were obtained at sufficiently high resolution (1.45 and 1.95 Å, respectively) and the corresponding structures were solved by molecular replacement. The crystal structures of the two enzymes (and two mutated variants) showed dimeric proteins. Complementary biophysical and molecular biology studies unveiled the diverse structural bases of the dimeric nature of the two enzymes. Intermolecular disulfide bridge and parallel association between two α-helices, among other interactions, were identified at the dimer interfaces. Interestingly, one of the r Cvi UPO variants incorporated the ability to produce fatty acid diepoxides-reactive compounds with valuable cross-linking capabilities-due to removal of the enzyme C-terminal tail located near the entrance of the heme access channel. In conclusion, different dimeric arrangements could be described in (short) UPO crystal structures.
Organizational Affiliation:
Centro de Investigaciones Biológicas "Margarita Salas" (CIB), CSIC, E-28040 Madrid, Spain.