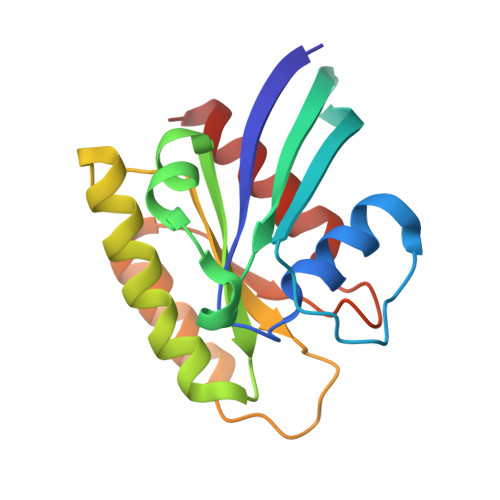
Far-reaching effects of tyrosine64 phosphorylation on Ras revealed with BeF 3 - complexes.
Baumann, P., Jin, Y.(2024) Commun Chem 7: 19-19
- PubMed: 38297137
- DOI: https://doi.org/10.1038/s42004-024-01105-6
- Primary Citation of Related Structures:
8BWG, 8CNJ, 8CNN - PubMed Abstract:
Tyrosine phosphorylation on Ras by Src kinase is known to uncouple Ras from upstream regulation and downstream communication. However, the mechanisms by which phosphorylation modulates these interactions have not been detailed. Here, the major mono-phosphorylation level on tyrosine64 is quantified by 31 P NMR and mutagenesis. Crystal structures of unphosphorylated and tyrosine64-phosphorylated Ras in complex with a BeF 3 - ground state analogue reveal "closed" Ras conformations very different from those of the "open" conformations previously observed for non-hydrolysable GTP analogue structures of Ras. They deliver new mechanistic and conformational insights into intrinsic GTP hydrolysis. Phosphorylation of tyrosine64 delivers conformational changes distant from the active site, showing why phosphorylated Ras has reduced affinity to its downstream effector Raf. 19 F NMR provides evidence for changes in the intrinsic GTPase and nucleotide exchange rate and identifies the concurrent presence of a major "closed" conformation alongside a minor yet functionally important "open" conformation at the ground state of Ras. This study expands the application of metal fluoride complexes in revealing major and minor conformational changes of dynamic and modified Ras proteins.
Organizational Affiliation:
School of Chemistry, Cardiff University, Park Place, Cardiff, CF10 3AT, UK.