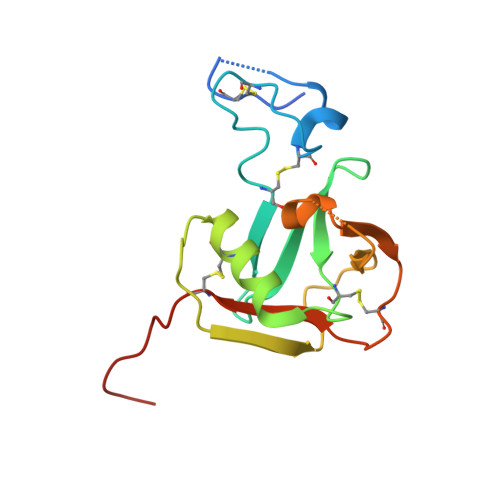
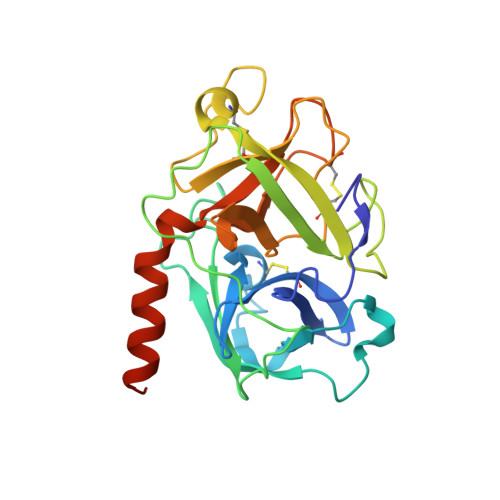
Structure-based discovery of dual pathway inhibitors for SARS-CoV-2 entry.
Wang, H., Yang, Q., Liu, X., Xu, Z., Shao, M., Li, D., Duan, Y., Tang, J., Yu, X., Zhang, Y., Hao, A., Wang, Y., Chen, J., Zhu, C., Guddat, L., Chen, H., Zhang, L., Chen, X., Jiang, B., Sun, L., Rao, Z., Yang, H.(2023) Nat Commun 14: 7574-7574
- PubMed: 37990007
- DOI: https://doi.org/10.1038/s41467-023-42527-5
- Primary Citation of Related Structures:
7XYD, 7Y0E, 7Y0F, 8HD8, 8HE9, 8HEI, 8HEN, 8HET, 8HFV - PubMed Abstract:
Since 2019, SARS-CoV-2 has evolved rapidly and gained resistance to multiple therapeutics targeting the virus. Development of host-directed antivirals offers broad-spectrum intervention against different variants of concern. Host proteases, TMPRSS2 and CTSL/CTSB cleave the SARS-CoV-2 spike to play a crucial role in the two alternative pathways of viral entry and are characterized as promising pharmacological targets. Here, we identify compounds that show potent inhibition of these proteases and determine their complex structures with their respective targets. Furthermore, we show that applying inhibitors simultaneously that block both entry pathways has a synergistic antiviral effect. Notably, we devise a bispecific compound, 212-148, exhibiting the dual-inhibition ability of both TMPRSS2 and CTSL/CTSB, and demonstrate antiviral activity against various SARS-CoV-2 variants with different viral entry profiles. Our findings offer an alternative approach for the discovery of SARS-CoV-2 antivirals, as well as application for broad-spectrum treatment of viral pathogenic infections with similar entry pathways.
Organizational Affiliation:
Shanghai Institute for Advanced Immunochemical Studies and School of Life Science and Technology, ShanghaiTech University, Shanghai, China.