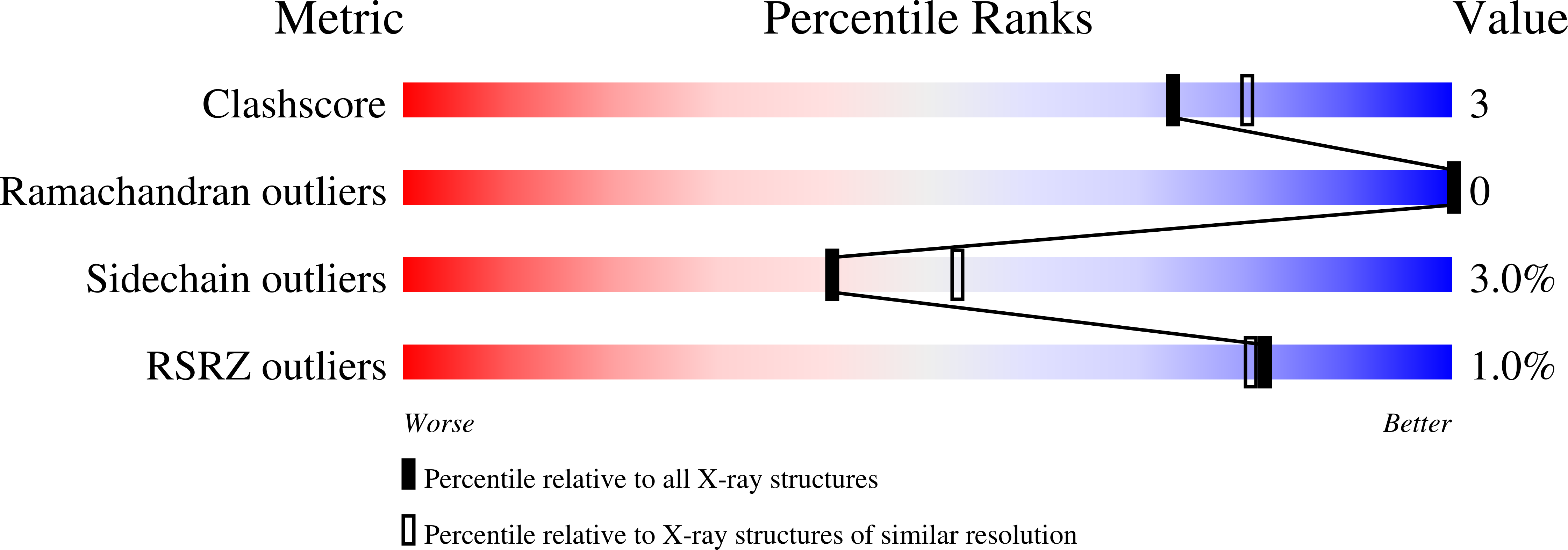
Oligosaccharide specificity of a family 7 endoglucanase: insertion of potential sugar-binding subsites.
Davies, G.J., Ducros, V., Lewis, R.J., Borchert, T.V., Schulein, M.(1997) J Biotechnol 57: 91-100
- PubMed: 9335168
- DOI: https://doi.org/10.1016/s0168-1656(97)00092-8
- Primary Citation of Related Structures:
1A39 - PubMed Abstract:
Family 7 of the glycosyl hydrolases contains both endoglucanases and cellobiohydrolases. In addition to their different catalytic activities on crystalline substrates, the cellobiohydrolases differ from the endoglucanases in their activity on longer soluble substrates, indicative of a greater number of subsites on the enzyme. A double mutant (S37W, P39W) of the Humicola insolens endoglucanase I (EG I) has been constructed in order to mimic aspects of the subsite structure of the corresponding family 7 cellobiohydrolase, cellobiohydrolase-I (CBH I). The 3-D crystal structure of the double mutant has been solved and refined to a crystallographic R-factor of 0.17 at a resolution of 2.2 A (1 A = 0.1 nm). The two mutant tryptophans are clearly visible in the electron density and are in the same orientation as those found in the substrate binding groove of CBH I. In addition to the substitutions, the C-terminal amino acids (399QELQ), disordered in the native enzyme structure, are clearly visible and there are a small number of minor loop movements associated with differences in crystal packing. Kinetic determinations show that the S37W, P39W mutant EG I has almost identical activity, compared to native EG I, on small soluble cellodextrins. On phosphoric acid swollen cellulose there is a small (30%), but significant, decrease in the apparent KM indicating that the double mutant may indeed exhibit stronger binding to longer polymeric substrates.
Organizational Affiliation:
Department of Chemistry, University of York, Heslington, UK. [email protected]