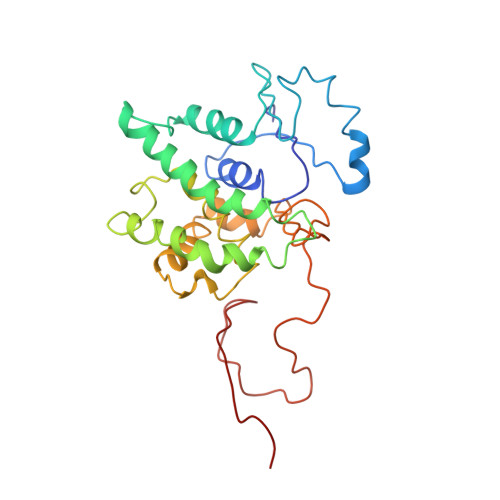
Structure of the ankyrin-binding domain of alpha-Na,K-ATPase.
Zhang, Z., Devarajan, P., Dorfman, A.L., Morrow, J.S.(1998) J Biol Chem 273: 18681-18684
- PubMed: 9668035
- DOI: https://doi.org/10.1074/jbc.273.30.18681
- Primary Citation of Related Structures:
1BG5 - PubMed Abstract:
The ankyrin 33-residue repeating motif, an L-shaped structure with protruding beta-hairpin tips, mediates specific macromolecular interactions with cytoskeletal, membrane, and regulatory proteins. The association between ankyrin and alpha-Na,K-ATPase, a ubiquitous membrane protein critical to vectorial transport of ions and nutrients, is required to assemble and stabilize Na,K-ATPase at the plasma membrane. alpha-Na,K-ATPase binds both red cell ankyrin (AnkR, a product of the ANK1 gene) and Madin-Darby canine kidney cell ankyrin (AnkG, a product of the ANK3 gene) utilizing residues 142-166 (SYYQEAKSSKIMESFK NMVPQQALV) in its second cytoplasmic domain. Fusion peptides of glutathione S-transferase incorporating these 25 amino acids bind specifically to purified ankyrin (Kd = 118 +/- 50 nM). The three-dimensional structure (2.6 A) of this minimal ankyrin-binding motif, crystallized as the fusion protein, reveals a 7-residue loop with one charged hydrophilic face capping a double beta-strand. Comparison with ankyrin-binding sequences in p53, CD44, neurofascin/L1, and the inositol 1,4,5-trisphosphate receptor suggests that the valency and specificity of ankyrin binding is achieved by the interaction of 5-7-residue surface loops with the beta-hairpin tips of multiple ankyrin repeat units.
Organizational Affiliation:
Department of Pathology, Yale University School of Medicine, New Haven, Connecticut 06510, USA.