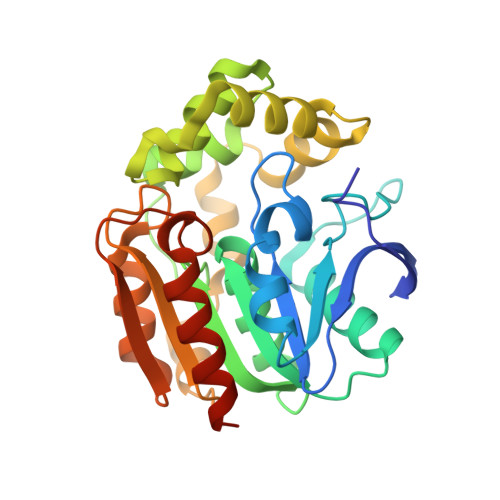
Crystal structures of a meta-cleavage product hydrolase from Pseudomonas fluorescens IP01 (CumD) complexed with cleavage products
Fushinobu, S., Saku, T., Hidaka, M., Jun, S.-Y., Nojiri, H., Yamane, H., Shoun, H., Omori, T., Wakagi, T.(2002) Protein Sci 11: 2184-2195
- PubMed: 12192074
- DOI: https://doi.org/10.1110/ps.0209602
- Primary Citation of Related Structures:
1IUN, 1IUO, 1IUP - PubMed Abstract:
2-Hydroxy-6-oxo-7-methylocta-2,4-dienoate hydrolase (CumD) from Pseudomonas fluorescens IP01 hydrolyzes a meta-cleavage product generated in the cumene (isopropylbenzene) degradation pathway. The crystal structures of the inactive S103A mutant of the CumD enzyme complexed with isobutyrate and acetate ions were determined at 1.6 and 2.0 A resolution, respectively. The isobutyrate and acetate ions were located at the same position in the active site, and occupied the site for a part of the hydrolysis product with CumD, which has the key determinant group for the substrate specificity of related hydrolases. One of the oxygen atoms of the carboxyl group of the isobutyrate ion was hydrogen bonded with a water molecule and His252. Another oxygen atom of the carboxyl group was situated in an oxyanion hole formed by the two main-chain N atoms. The isopropyl group of the isobutyric acid was recognized by the side-chains of the hydrophobic residues. The substrate-binding pocket of CumD was long, and the inhibition constants of various organic acids corresponded well to it. In comparison with the structure of BphD from Rhodococcus sp. RHA1, the structural basis for the substrate specificity of related hydrolases, is revealed.
Organizational Affiliation:
Department of Biotechnology, The University of Tokyo, Bunkyo-ku, Tokyo 113-8657, Japan. [email protected]