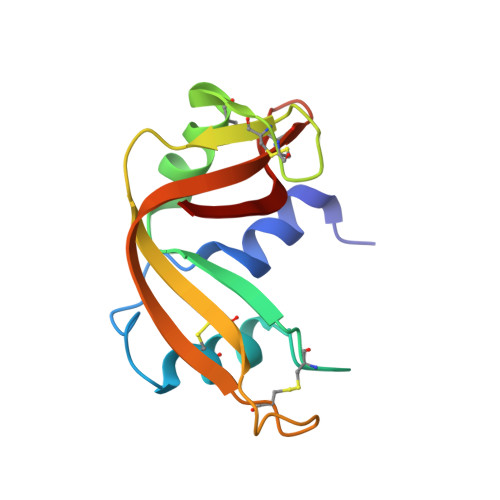
Minimization of cavity size ensures protein stability and folding: structures of Phe46-replaced bovine pancreatic RNase A
Kadonosono, T., Chatani, E., Hayashi, R., Moriyama, H., Ueki, T.(2003) Biochemistry 42: 10651-10658
- PubMed: 12962489
- DOI: https://doi.org/10.1021/bi034499w
- Primary Citation of Related Structures:
1IZP, 1IZQ, 1IZR - PubMed Abstract:
The Phe46 residue, located in the hydrophobic core of RNase A, was replaced with other hydrophobic residues, leucine, valine, or alanine, and their X-ray crystallographic structures were determined up to 1.50-1.80 A resolution in an attempt to examine the relationship between structural changes and conformational stability or folding kinetics. The backbone structure of F46L, F46V, and F46A was indistinguishable from that of the wild-type enzyme, retaining the correct active site structure. However, one water molecule was included in the hydrophobic core of F46A, forming two hydrogen bonds with the backbone peptide chain. The side chain of Met29 in F46V and F46A adopted two different conformations in an equal occupancy. A trapped water molecule and two conformations of Met29 represent changes that minimize the cavity volume. Nevertheless, the replacement of Phe46 with the above residues resulted in a marked decrease in both thermal stability and folding reaction. Thus, Phe46 ensures the thermal stability and the rapid and correct folding of RNase A by the role it plays in forming a highly packed, hydrophobic core.
Organizational Affiliation:
Division of Applied Life Sciences, Graduate School of Agriculture, Kyoto University, Sakyo, Kyoto 606-8502, Japan.