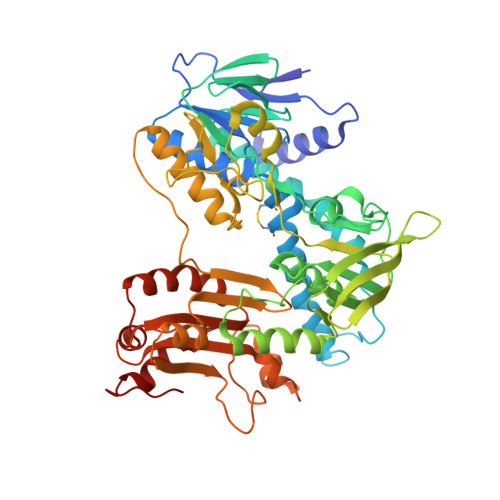
Structural Basis for CO2 Fixation by a Novel Member of the Disulfide Oxidoreductase Family of Enzymes, 2-Ketopropyl Coenzyme M Oxidoreductase/Carboxylase
Nocek, B., Jang, S.B., Jeong, M.S., Clark, D.D., Ensign, S.A., Peters, J.W.(2002) Biochemistry 41: 12907-12913
- PubMed: 12390015
- DOI: https://doi.org/10.1021/bi026580p
- Primary Citation of Related Structures:
1MO9, 1MOK - PubMed Abstract:
The NADPH:2-ketopropyl-coenzyme M oxidoreductase/carboxylase (2-KPCC) is the terminal enzyme in a metabolic pathway that results in the conversion of propylene to the central metabolite acetoacetate in Xanthobacter autotrophicus Py2. This enzyme is an FAD-containing enzyme that is a member of the NADPH:disulfide oxidoreductase (DSOR) family of enzymes that include glutathione reductase, dihydrolipoamide dehydrogenase, trypanothione reductase, thioredoxin reductase, and mercuric reductase. In contrast to the prototypical reactions catalyzed by members of the DSOR family, the NADPH:2-ketopropyl-coenzyme M oxidoreductase/carboxylase catalyzes the reductive cleavage of the thioether linkage of 2-ketopropyl-coenzyme M, and the subsequent carboxylation of the ketopropyl cleavage product, yielding the products acetoacetate and free coenzyme M. The structure of 2-KPCC reveals a unique active site in comparison to those of other members of the DSOR family of enzymes and demonstrates how the enzyme architecture has been adapted for the more sophisticated biochemical reaction. In addition, comparison of the structures in the native state and in the presence of bound substrate indicates the binding of the substrate 2-ketopropyl-coenzyme M induces a conformational change resulting in the collapse of the substrate access channel. The encapsulation of the substrate in this manner is reminiscent of the conformational changes observed in the well-characterized CO2-fixing enzyme ribulose 1,5-bisphosphate carboxylase/oxidase (Rubisco).
Organizational Affiliation:
Department of Chemistry and Biochemistry, Montana State University, Bozeman, Montana 59717, USA.