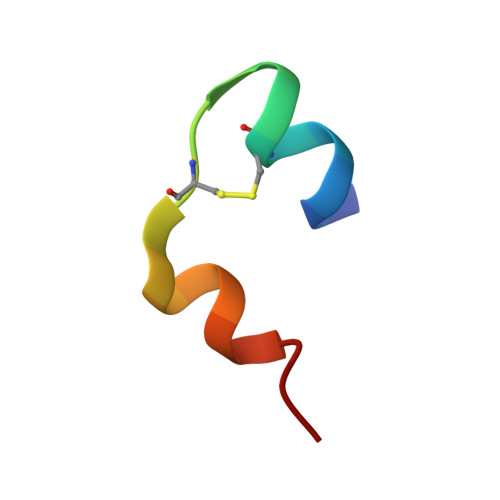
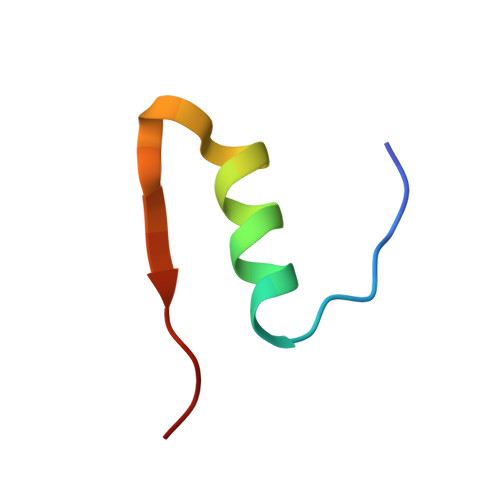
The structure of T6 human insulin at 1.0 A resolution.
Smith, G.D., Pangborn, W.A., Blessing, R.H.(2003) Acta Crystallogr D Biol Crystallogr 59: 474-482
- PubMed: 12595704
- DOI: https://doi.org/10.1107/s0907444902023685
- Primary Citation of Related Structures:
1MSO - PubMed Abstract:
The structure of T(6) human insulin has been determined at 120 K at a resolution of 1.0 A and refined to a residual of 0.183. As a result of cryofreezing, the first four residues of the B chain in one of the two crystallographically independent AB monomers in the hexameric [Zn(1/3)(AB)(2)Zn(1/3)](3) complex undergo a conformational shift that displaces the C(alpha) atom of PheB1 by 7.86 A relative to the room-temperature structure. A least-squares superposition of all backbone atoms of the room-temperature and low-temperature structures yielded a mean displacement of 0.422 A. Omitting the first four residues of the B chain reduced the mean displacement to 0.272 A. At 120 K, nine residues were found to exhibit two discrete side-chain conformations, but only two of these residues are in common with the seven residues found to have disordered side chains in the room-temperature structure. As a result of freezing, the disorder observed at room temperature in both ArgB22 side chains is eliminated. The close contact between pairs of O( epsilon 2) atoms in GluB13 observed at room temperature is maintained at cryotemperature and suggests that a carboxylate-carboxylic acid centered hydrogen bond exists [-C(=O)-O.H.O-C(=O)-] such that the H atom is equally shared between the two partially charged O atoms.
Organizational Affiliation:
Structural Biology and Biochemistry, Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada. [email protected]