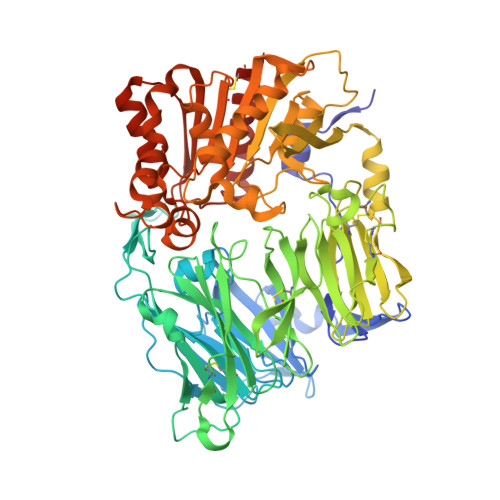
Crystal structure of human dipeptidyl peptidase IV/CD26 in complex with a substrate analogue
Rasmussen, H.B., Branner, S., Wiberg, F.C., Wagtmann, N.R.(2003) Nat Struct Biol 10: 19-25
- PubMed: 12483204
- DOI: https://doi.org/10.1038/nsb882
- Primary Citation of Related Structures:
1N1M - PubMed Abstract:
Dipeptidyl peptidase IV (DPP-IV/CD26) is a multifunctional type II transmembrane serine peptidase. This enzyme contributes to the regulation of various physiological processes, including blood sugar homeostasis, by cleaving peptide hormones, chemokines and neuropeptides. We have determined the 2.5 A structure of the extracellular region of DPP-IV in complex with the inhibitor valine-pyrrolidide. The catalytic site is located in a large cavity formed between the alpha/beta-hydrolase domain and an eight-bladed beta-propeller domain. Both domains participate in inhibitor binding. The structure indicates how substrate specificity is achieved and reveals a new and unexpected opening to the active site.
Organizational Affiliation:
Protein Chemistry, Research and Development, Novo Nordisk A/S, Novo Allé, DK-2880 Bagsvaerd, Denmark. [email protected]