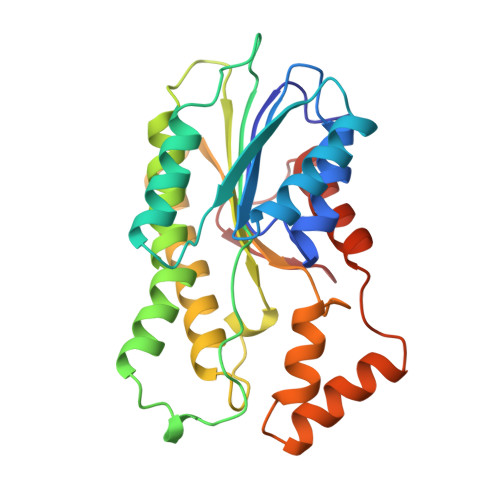
The 1.25 A crystal structure of sepiapterin reductase reveals its binding mode to pterins and brain neurotransmitters.
Auerbach, G., Herrmann, A., Gutlich, M., Fischer, M., Jacob, U., Bacher, A., Huber, R.(1997) EMBO J 16: 7219-7230
- PubMed: 9405351
- DOI: https://doi.org/10.1093/emboj/16.24.7219
- Primary Citation of Related Structures:
1NAS, 1OAA, 1SEP - PubMed Abstract:
Sepiapterin reductase catalyses the last steps in the biosynthesis of tetrahydrobiopterin, the essential co-factor of aromatic amino acid hydroxylases and nitric oxide synthases. We have determined the crystal structure of mouse sepiapterin reductase by multiple isomorphous replacement at a resolution of 1.25 A in its ternary complex with oxaloacetate and NADP. The homodimeric structure reveals a single-domain alpha/beta-fold with a central four-helix bundle connecting two seven-stranded parallel beta-sheets, each sandwiched between two arrays of three helices. Ternary complexes with the substrate sepiapterin or the product tetrahydrobiopterin were studied. Each subunit contains a specific aspartate anchor (Asp258) for pterin-substrates, which positions the substrate side chain C1'-carbonyl group near Tyr171 OH and NADP C4'N. The catalytic mechanism of SR appears to consist of a NADPH-dependent proton transfer from Tyr171 to the substrate C1' and C2' carbonyl functions accompanied by stereospecific side chain isomerization. Complex structures with the inhibitor N-acetyl serotonin show the indoleamine bound such that both reductase and isomerase activity for pterins is inhibited, but reaction with a variety of carbonyl compounds is possible. The complex structure with N-acetyl serotonin suggests the possibility for a highly specific feedback regulatory mechanism between the formation of indoleamines and pteridines in vivo.
Organizational Affiliation:
Max-Planck-Institut für Biochemie, Abt. Strukturforschung, Am Klopferspitz 18a, D-82152 Martinsried, Germany. [email protected]