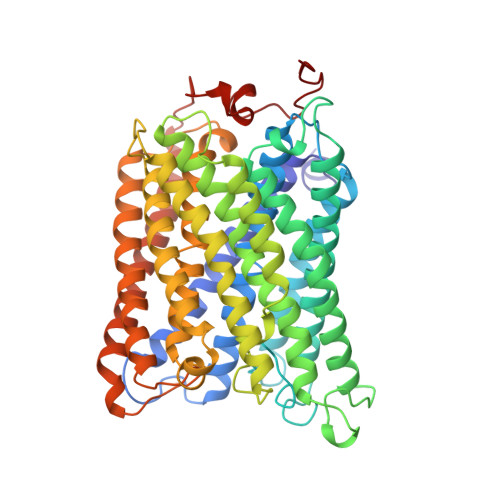
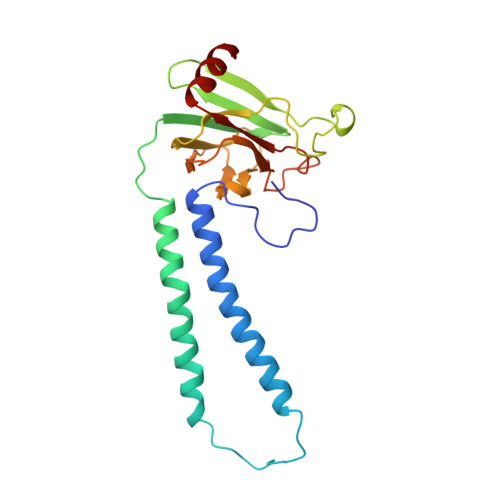
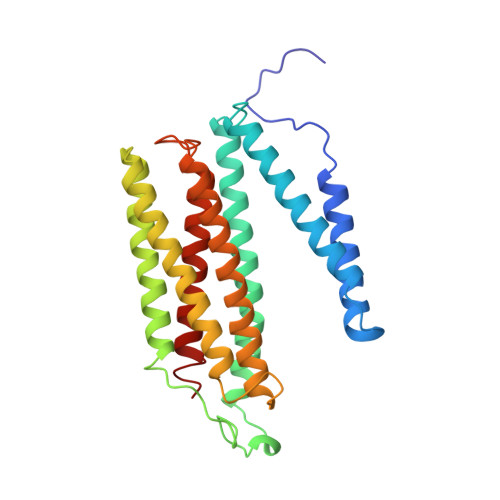
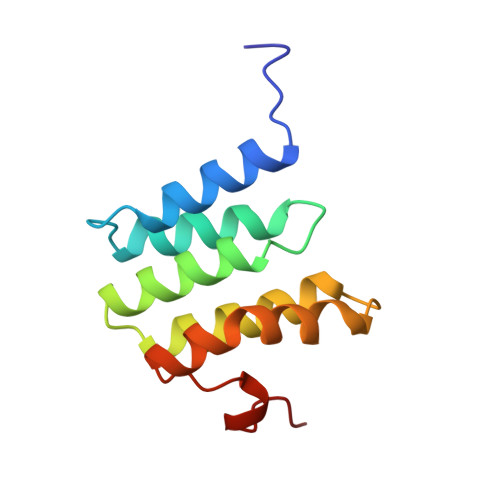
Redox-coupled crystal structural changes in bovine heart cytochrome c oxidase.
Yoshikawa, S., Shinzawa-Itoh, K., Nakashima, R., Yaono, R., Yamashita, E., Inoue, N., Yao, M., Fei, M.J., Libeu, C.P., Mizushima, T., Yamaguchi, H., Tomizaki, T., Tsukihara, T.(1998) Science 280: 1723-1729
- PubMed: 9624044
- DOI: https://doi.org/10.1126/science.280.5370.1723
- Primary Citation of Related Structures:
1OCO, 1OCR, 1OCZ, 2OCC - PubMed Abstract:
Crystal structures of bovine heart cytochrome c oxidase in the fully oxidized, fully reduced, azide-bound, and carbon monoxide-bound states were determined at 2.30, 2.35, 2.9, and 2.8 angstrom resolution, respectively. An aspartate residue apart from the O2 reduction site exchanges its effective accessibility to the matrix aqueous phase for one to the cytosolic phase concomitantly with a significant decrease in the pK of its carboxyl group, on reduction of the metal sites. The movement indicates the aspartate as the proton pumping site. A tyrosine acidified by a covalently linked imidazole nitrogen is a possible proton donor for the O2 reduction by the enzyme.
Organizational Affiliation:
Department of Life Science, Himeji Institute of Technology and CREST, Japan Science and Technology Corporation (JST), Kamigohri Akoh, Hyogo 678-1297, Japan.