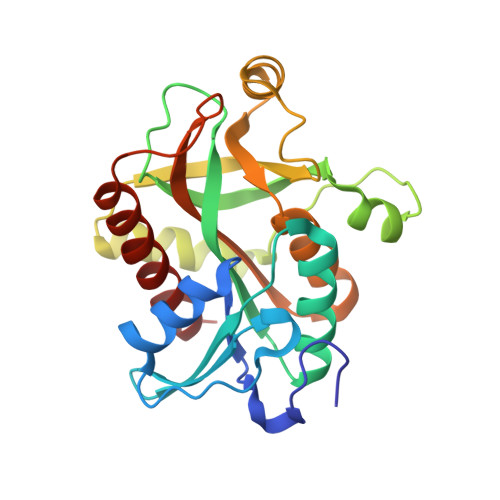
Crystal Structure of Purine Nucleoside Phosphorylase from Thermus Thermophilus
Tahirov, T.H., Inagaki, E., Ohshima, N., Kitao, T., Kuroishi, C., Ukita, Y., Takio, K., Kobayashi, M., Kuramitsu, S., Yokoyama, S., Miyano, M.(2004) J Mol Biol 337: 1149
- PubMed: 15046984
- DOI: https://doi.org/10.1016/j.jmb.2004.02.016
- Primary Citation of Related Structures:
1ODI, 1ODJ, 1ODK, 1ODL - PubMed Abstract:
The purine nucleoside phosphorylase from Thermus thermophilus crystallized in space group P4(3)2(1)2 with the unit cell dimensions a = 131.9 A and c = 169.9 A and one biologically active hexamer in the asymmetric unit. The structure was solved by the molecular replacement method and refined at a 1.9A resolution to an r(free) value of 20.8%. The crystals of the binary complex with sulfate ion and ternary complexes with sulfate and adenosine or guanosine were also prepared and their crystal structures were refined at 2.1A, 2.4A and 2.4A, respectively. The overall structure of the T.thermophilus enzyme is similar to the structures of hexameric enzymes from Escherichia coli and Sulfolobus solfataricus, but significant differences are observed in the purine base recognition site. A base recognizing aspartic acid, which is conserved among the hexameric purine nucleoside phosphorylases, is Asn204 in the T.thermophilus enzyme, which is reminiscent of the base recognizing asparagine in trimeric purine nucleoside phosphorylases. Isothermal titration calorimetry measurements indicate that both adenosine and guanosine bind the enzyme with nearly similar affinity. However, the functional assays show that as in trimeric PNPs, only the guanosine is a true substrate of the T.thermophilus enzyme. In the case of adenosine recognition, the Asn204 forms hydrogen bonds with N6 and N7 of the base. While in the case of guanosine recognition, the Asn204 is slightly shifted together with the beta(9)alpha(7) loop and predisposed to hydrogen bond formation with O6 of the base in the transition state. The obtained experimental data suggest that the catalytic properties of the T.thermophilus enzyme are reminiscent of the trimeric rather than hexameric purine nucleoside phosphorylases.
Organizational Affiliation:
Highthroughput Factory, RIKEN Harima Institute, 1-1-1 Kouto, Mikazuki-cho, Sayo-gun, Hyogo 679-5148, Japan. [email protected]