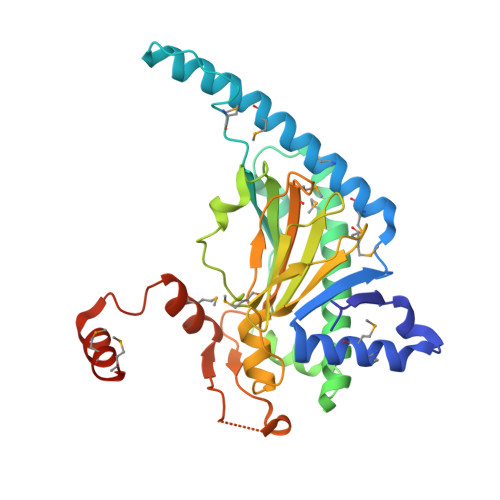
Crystal Structure and Mechanistic Implications of 1-Aminocyclopropane-1-Carboxylic Acid Oxidase (the Ethyling Forming Enzyme)
Zhang, Z., Ren, J.-S., Clifton, I.J., Schofield, C.J.(2004) Chem Biol 11: 1383
- PubMed: 15489165
- DOI: https://doi.org/10.1016/j.chembiol.2004.08.012
- Primary Citation of Related Structures:
1W9Y, 1WA6 - PubMed Abstract:
The final step in the biosynthesis of the plant signaling molecule ethylene is catalyzed by 1-aminocyclopropane-1-carboxylic acid oxidase (ACCO). ACCO requires bicarbonate as an activator and catalyzes the oxidation of ACC to give ethylene, CO2, and HCN. We report crystal structures of ACCO in apo-form (2.1 A resolution) and complexed with Fe(II) (2.55 A) or Co(II) (2.4 A). The active site contains a single Fe(II) ligated by three residues (His177, Asp179, and His234), and it is relatively open compared to those of the 2-oxoglutarate oxygenases. The side chains of Arg175 and Arg244, proposed to be involved in binding bicarbonate, project away from the active site, but conformational changes may allow either or both to enter the active site. The structures will form a basis for future mechanistic and inhibition studies.
Organizational Affiliation:
The Oxford Centre for Molecular Sciences, The Department of Chemistry, University of Oxford, Chemistry Research Laboratory, Mansfield Road, Oxford OX1 3TA, United Kingdom.