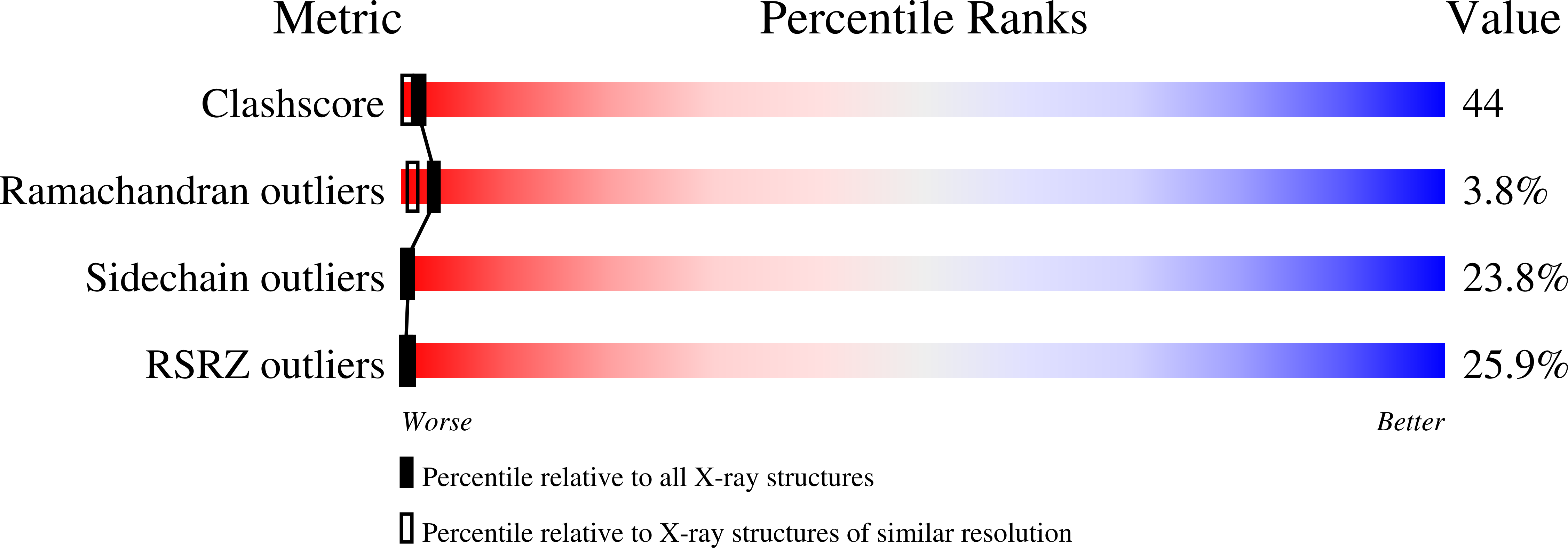Structure of a Pseudomerohedrally Twinned Monoclinic Crystal Form of a Pyridoxal Phosphate-Dependent Catalytic Antibody
Golinelli-Pimpaneau, B.(2005) Acta Crystallogr D Biol Crystallogr 61: 472
- PubMed: 15805602
- DOI: https://doi.org/10.1107/S0907444905003331
- Primary Citation of Related Structures:
1WC7 - PubMed Abstract:
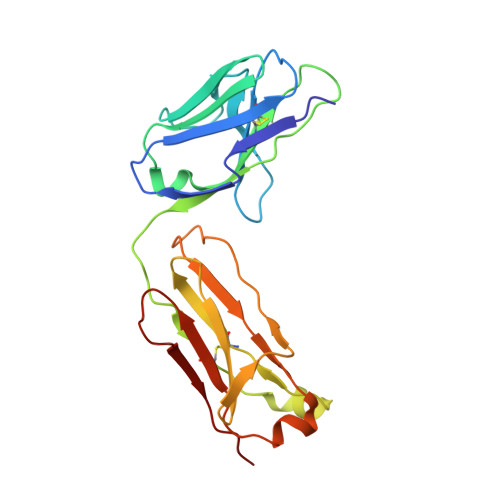
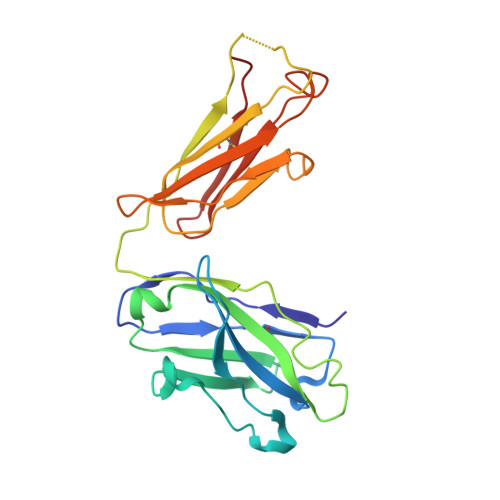
The purification, crystallization and structure determination at 2.3 A resolution of the complex of the pyridoxal-5'-phosphate (PLP) dependent catalytic antibody 15A9 with a phosphopyridoxyl-L-alanine (PPL-L-alanine) substrate analogue are described. The crystal belongs to space group P2(1), with two molecules in the asymmetric unit related by non-crystallographic symmetry. The unit-cell parameters are a = 63.5, b = 81.7, c = 79.3 A and beta is fortuitously 90 degrees . Refinement of the structure converged at unacceptably high R factors. Although the traditional analysis of intensity distribution did not indicate twinning, pseudomerohedral twinning was revealed by a newer test based on local intensity differences [Padilla & Yeates (2003), Acta Cryst. D59, 1124-1130]. When the potential twinning operator was included in SHELX, the structure could be satisfactorily refined with a twinning fraction of 0.46, indicating a nearly perfect hemihedrally twinned crystal. One of the active sites is occupied by the phosphopyridoxyl-L-alanine ligand, while one iodide ion mimics the cofactor phosphate group in the other. Four other iodide ions are present in the structure: two are involved in specific intermolecular contacts and two dictate the conformation of the CDRH3 loop in each molecule.
Organizational Affiliation:
Laboratoire d'Enzymologie et de Biochimie Structurales, Bâtiment 34, CNRS, 1 Avenue de la Terrasse, 91190 Gif-sur-Yvette, France. [email protected]