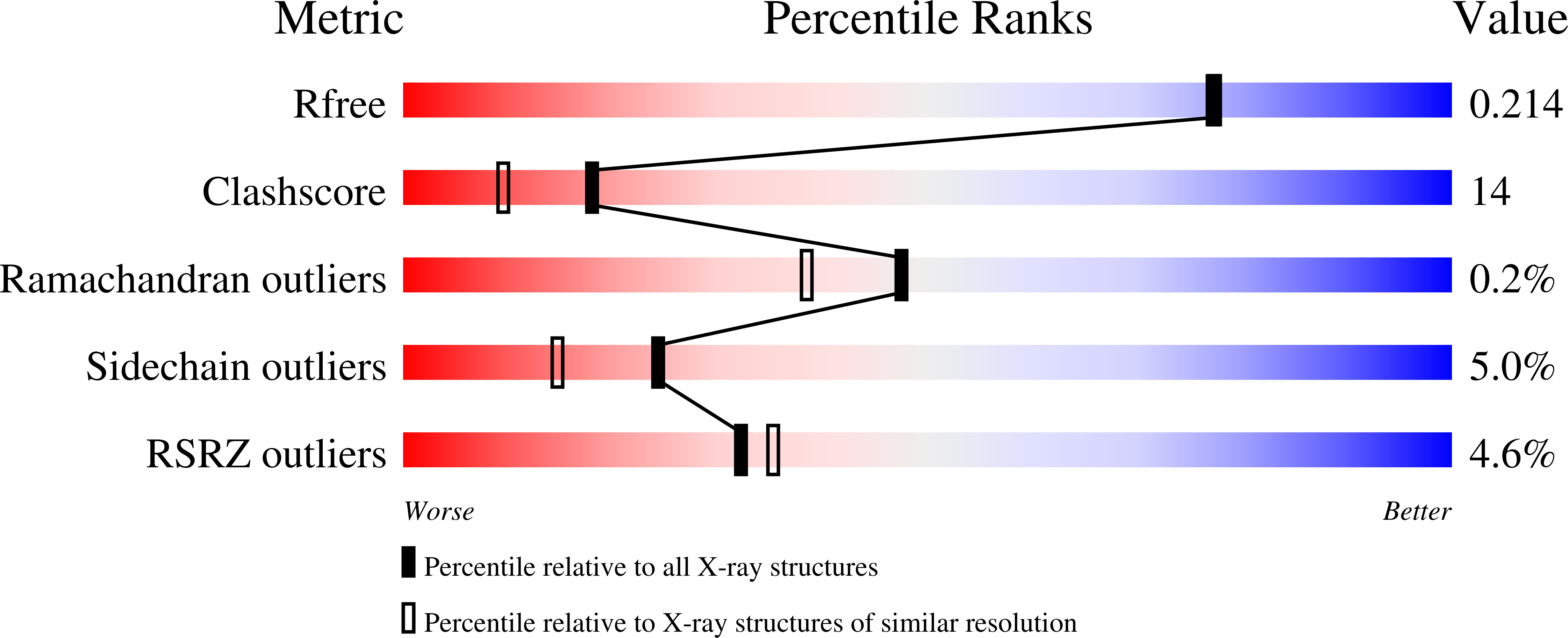
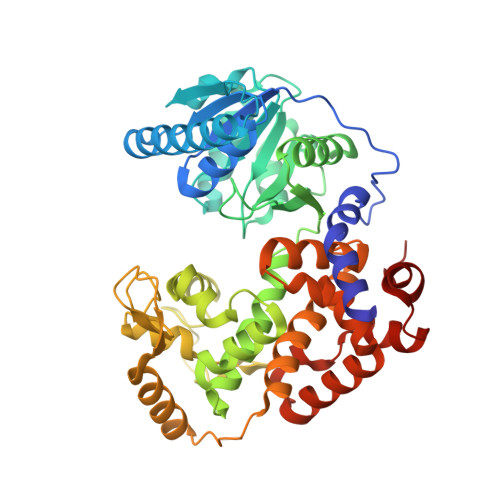
Structural Studies of MFE-1: the 1.9A Crystal Structure of the Dehydrogenase Part of Rat Peroxisomal MFE-1
Taskinen, J.P., Kiema, T.R., Hiltunen, J.K., Wierenga, R.K.(2006) J Mol Biol 355: 734-746
- PubMed: 16330050
- DOI: https://doi.org/10.1016/j.jmb.2005.10.085
- Primary Citation of Related Structures:
1ZCJ - PubMed Abstract:
The 1.9 A structure of the C-terminal dehydrogenase part of the rat peroxisomal monomeric multifunctional enzyme type 1 (MFE-1) has been determined. In this construct (residues 260-722 and referred to as MFE1-DH) the N-terminal hydratase part of MFE-1 has been deleted. The structure of MFE1-DH shows that it consists of an N-terminal helix, followed by a Rossmann-fold domain (domain C), followed by two tightly associated helical domains (domains D and E), which have similar topology. The structure of MFE1-DH is compared with the two known homologous structures: human mitochondrial 3-hydroxyacyl-CoA dehydrogenase (HAD; sequence identity is 33%) (which is dimeric and monofunctional) and with the dimeric multifunctional alpha-chain (alphaFOM; sequence identity is 28%) of the bacterial fatty acid beta-oxidation alpha2beta2-multienzyme complex. Like MFE-1, alphaFOM has an N-terminal hydratase part and a C-terminal dehydrogenase part, and the structure comparisons show that the N-terminal helix of MFE1-DH corresponds to the alphaFOM linker helix, located between its hydratase and dehydrogenase part. It is also shown that this helix corresponds to the C-terminal helix-10 of the hydratase/isomerase superfamily, suggesting that functionally it belongs to the N-terminal hydratase part of MFE-1.
Organizational Affiliation:
Biocenter Oulu and Department of Biochemistry, University of Oulu, P.O. Box 3000, FIN-90014, Finland.