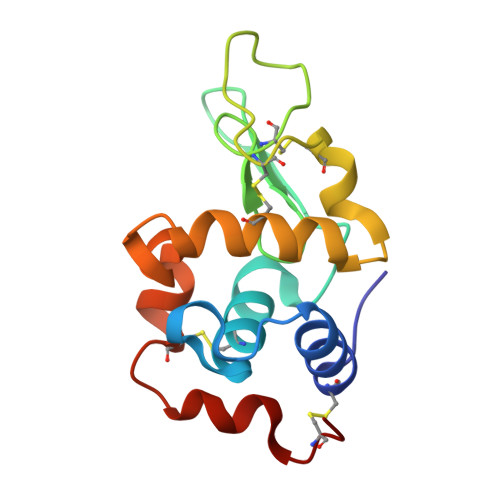
Chemical Basis for the Affinity Maturation of a Camel Single Domain Antibody
De Genst, E., Handelberg, F., Van Meirhaeghe, A., Vynck, S., Loris, R., Wyns, L., Muyldermans, S.(2004) J Biol Chem 279: 53593-53601
- PubMed: 15383540
- DOI: https://doi.org/10.1074/jbc.M407843200
- Primary Citation of Related Structures:
1XFP - PubMed Abstract:
Affinity maturation of classic antibodies supposedly proceeds through the pre-organization of the reactive germ line conformational isomer. It is less evident to foresee how this can be accomplished by camelid heavy-chain antibodies lacking light chains. Although these antibodies are subjected to somatic hypermutation, their antigen-binding fragment consists of a single domain with restricted flexibility in favor of binding energy. An antigen-binding domain derived from a dromedary heavy-chain antibody, cAb-Lys3, accumulated five amino acid substitutions in CDR1 and CDR2 upon maturation against lysozyme. Three of these residues have hydrophobic side chains, replacing serines, and participate in the hydrophobic core of the CDR1 in the mature antibody, suggesting that conformational rearrangements might occur in this loop during maturation. However, transition state analysis of the binding kinetics of mature cAb-Lys3 and germ line variants show that the maturation of this antibody relies on events late in the reaction pathway. This is reflected by a limited perturbation of k(a) and a significantly decreased k(d) upon maturation. In addition, binding reactions and the maturation event are predominantly enthalpically driven. Therefore, maturation proceeds through the increase of favorable binding interactions, or by the reduction of the enthalpic penalty for desolvation, as opposed to large entropic penalties associated with conformational changes and structural plasticity. Furthermore, the crystal structure of the mutant with a restored germ line CDR2 sequence illustrates that the matured hydrophobic core of CDR1 in cAb-Lys3 might be compensated in the germ line precursor by burying solvent molecules engaged in a stable hydrogen-bonding network with CDR1 and CDR2.
Organizational Affiliation:
Department of Molecular and Cellular Interactions, Vlaams Interuniversitair Instituut voor Biotechnologie, Vrije Universiteit Brussel, Pleinlaan 2, B-1050 Brussels, Belgium. [email protected]