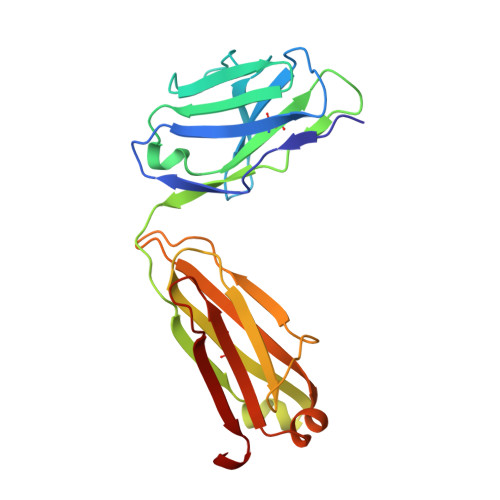
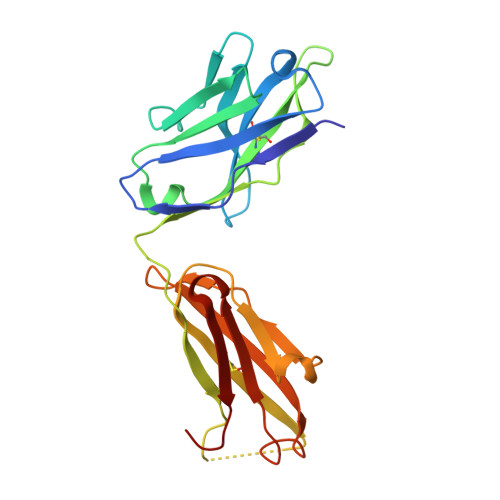
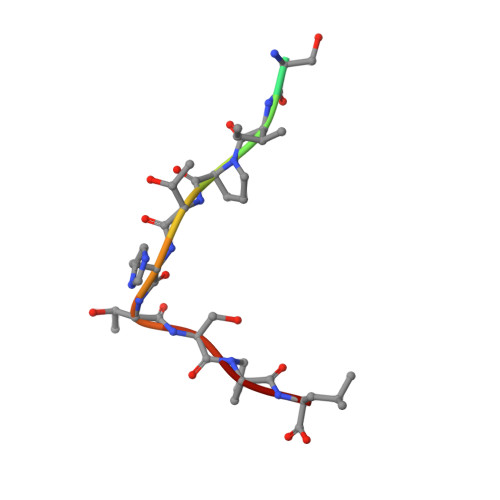
Differential epitope positioning within the germline antibody paratope enhances promiscuity in the primary immune response.
Sethi, D.K., Agarwal, A., Manivel, V., Rao, K.V., Salunke, D.M.(2006) Immunity 24: 429-438
- PubMed: 16618601
- DOI: https://doi.org/10.1016/j.immuni.2006.02.010
- Primary Citation of Related Structures:
2A6D, 2A6I, 2A6J, 2A6K - PubMed Abstract:
Correlation between the promiscuity of the primary antibody response and conformational flexibility in a germline antibody was addressed by using germline antibody 36-65. Crystallographic analyses of the 36-65 Fab with three independent dodecapeptides provided mechanistic insights into the generation of antibody diversity. While four antigen-free Fab molecules provided a quantitative description of the conformational repertoire of the antibody CDRs, three Fab molecules bound to structurally diverse peptide epitopes exhibited a common paratope conformation. Each peptide revealed spatially different footprints within the antigen-combining site. However, a conformation-specific lock involving two shared residues, which were also associated with hapten binding, was discernible. Unlike the hapten, the peptides interacted with residues that undergo somatic mutations, suggesting a possible mechanism for excluding "nonspecific" antigens during affinity maturation. The observed multiple binding modes of diverse epitopes within a common paratope conformation of a germline antibody reveal a simple, yet elegant, mechanism for expanding the primary antibody repertoire.
Organizational Affiliation:
National Institute of Immunology, New Delhi 110067, India.