Flexibility of Thiamine Diphosphate Revealed by Kinetic Crystallographic Studies of the Reaction of Pyruvate-Ferredoxin Oxidoreductase with Pyruvate.
Cavazza, C., Contreras-Martel, C., Pieulle, L., Chabriere, E., Hatchikian, E.C., Fontecilla-Camps, J.C.(2006) Structure 14: 217
- PubMed: 16472741
- DOI: https://doi.org/10.1016/j.str.2005.10.013
- Primary Citation of Related Structures:
2C3M, 2C3O, 2C3P, 2C3U, 2C3Y, 2C42, 2UZA - PubMed Abstract:
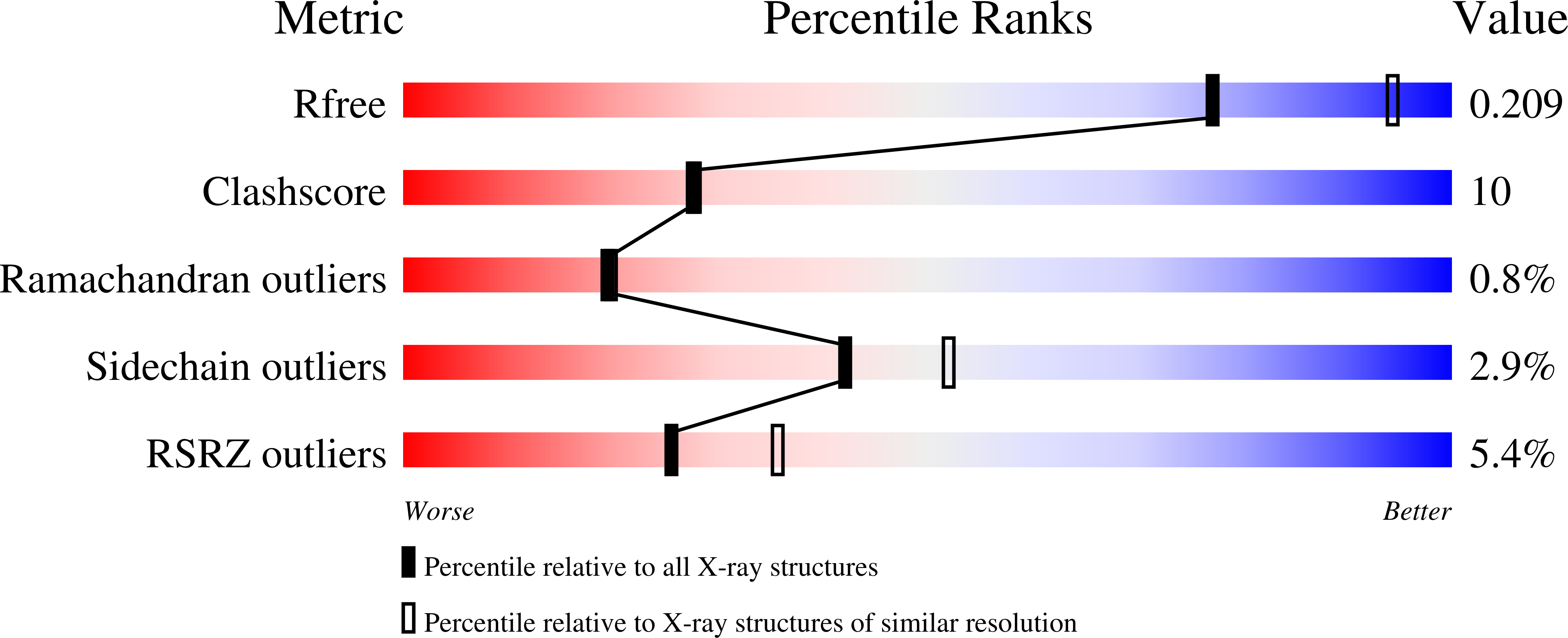
Pyruvate-ferredoxin oxidoreductases (PFOR) are unique among thiamine pyrophosphate (ThDP)-containing enzymes in giving rise to a rather stable cofactor-based free-radical species upon the decarboxylation of their first substrate, pyruvate. We have obtained snapshots of unreacted and partially reacted (probably as a tetrahedral intermediate) pyruvate-PFOR complexes at different time intervals. We conclude that pyruvate decarboxylation involves very limited substrate-to-product movements but a significant displacement of the thiazolium moiety of ThDP. In this respect, PFOR seems to differ substantially from other ThDP-containing enzymes, such as transketolase and pyruvate decarboxylase. In addition, exposure of PFOR to oxygen in the presence of pyruvate results in significant inhibition of catalytic activity, both in solution and in the crystals. Examination of the crystal structure of inhibited PFOR suggests that the loss of activity results from oxime formation at the 4' amino substituent of the pyrimidine moiety of ThDP.
Organizational Affiliation:
Laboratoire de Cristallographie et Cristallogenèse des Protéines, Institut de Biologie Structurale Jean-Pierre Ebel, CEA, UJF, CNRS, 41 rue Jules Horowitz, 38027 Grenoble Cedex 1, France.