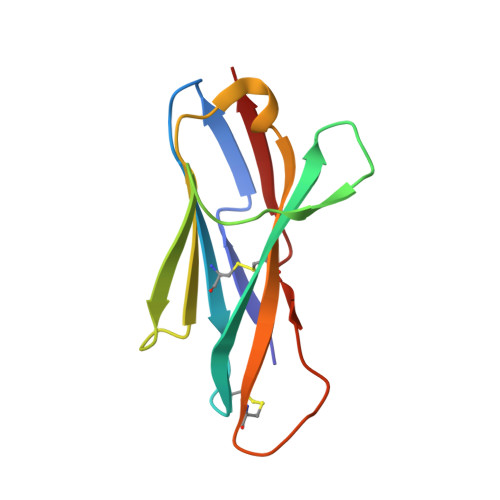
Structure of a shark IgNAR antibody variable domain and modeling of an early-developmental isotype.
Streltsov, V.A., Carmichael, J.A., Nuttall, S.D.(2005) Protein Sci 14: 2901-2909
- PubMed: 16199666
- DOI: https://doi.org/10.1110/ps.051709505
- Primary Citation of Related Structures:
2COQ - PubMed Abstract:
The new antigen receptor (IgNAR) antibodies from sharks are disulphide bonded dimers of two protein chains, each containing one variable and five constant domains. Three types of IgNAR variable domains have been discovered, with Type 3 appearing early in shark development and being overtaken by the antigen-driven affinity-matured Type 1 and 2 response. Here, we have determined the first structure of a naturally occurring Type 2 IgNAR variable domain, and identified the disulphide bond that links and stabilizes the CDR1 and CDR3 loops. This disulphide bridge locks the CDR3 loop in an "upright" conformation in contrast to other shark antibody structures, where a more lateral configuration is observed. Further, we sought to model the Type 3 isotype based on the crystallographic structure reported here. This modeling indicates (1) that internal Type 3-specific residues combine to pack into a compact immunoglobulin core that supports the CDR loop regions, and (2) that despite apparent low-sequence variability, there is sufficient plasticity in the CDR3 loop to form a conformationally diverse antigen-binding surface.
Organizational Affiliation:
CSIRO Molecular and Health Technologies, 343 Royal Parade, Parkville, Victoria 3052, Australia. [email protected]