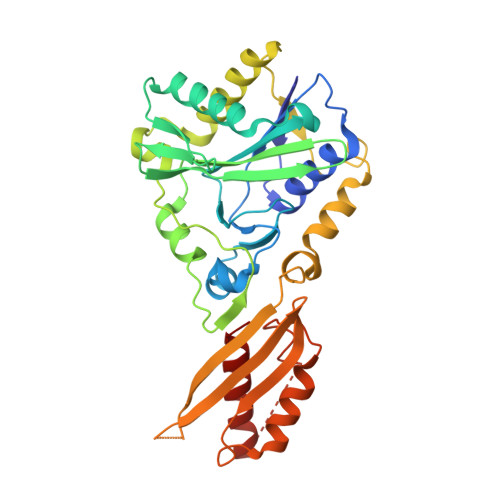
Crystal structure of bovine Lipoyltransferase in complex with lipoyl-AMP
Fujiwara, K., Hosaka, H., Matsuda, M., Okamura-Ikeda, K., Motokawa, Y., Suzuki, M., Nakagawa, A., Taniguchi, H.(2007) J Mol Biol 371: 222-234
- PubMed: 17570395
- DOI: https://doi.org/10.1016/j.jmb.2007.05.059
- Primary Citation of Related Structures:
2E5A - PubMed Abstract:
Lipoic acid is an essential cofactor of the alpha-ketoacid dehydrogenase complexes and the glycine cleavage system. It is covalently attached to a specific lysine residue of the subunit of the complexes. The bovine lipoyltransferase (bLT) catalyzes the lipoic acid attachment reaction using lipoyl-AMP as a substrate, forming a lipoylated protein and AMP. To gain insights into the reaction mechanism at the atomic level, we have determined the crystal structure of bLT at 2.10 A resolution. Unexpectedly, the purified recombinant bLT contains endogenous lipoyl-AMP. The structure of bLT consists of N-terminal and C-terminal domains, and lipoyl-AMP is bound to the active site in the N-terminal domain, adopting a U-shaped conformation. The lipoyl moiety is buried in the hydrophobic pocket, forming van der Waals interactions, and the AMP moiety forms numerous hydrogen bonds with bLT in another tunnel-like cavity. These interactions work together to expose the C10 atom of lipoyl-AMP to the surface of the bLT molecule. The carbonyl oxygen atom of lipoyl-AMP interacts with the invariant Lys135. The interaction might stimulate the positive charge of the C10 atom of lipoyl-AMP, and consequently facilitate the nucleophilic attack by the lysine residue of the lipoate-acceptor protein, accompanying the bond cleavage between the carbonyl group and the phosphate group. We discuss the structural differences between bLT and the lipoate-protein ligase A from Escherichia coli and Thermoplasma acidophilum. We further demonstrate that bLT in mitochondria also contains endogenous lipoylmononucleotide, being ready for the lipoylation of apoproteins.
Organizational Affiliation:
Institute for Enzyme Research, the University of Tokushima, Tokushima 770-8503, Japan. [email protected]