Structural Basis of the Inhibition of Golgi alpha-Mannosidase II by Mannostatin A and the Role of the Thiomethyl Moiety in Ligand-Protein Interactions.
Kawatkar, S.P., Kuntz, D.A., Woods, R.J., Rose, D.R., Boons, G.J.(2006) J Am Chem Soc 128: 8310-8319
- PubMed: 16787095
- DOI: https://doi.org/10.1021/ja061216p
- Primary Citation of Related Structures:
2F7O, 2F7P - PubMed Abstract:
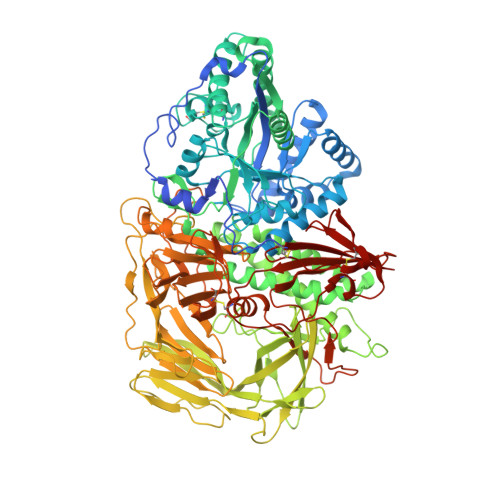
The X-ray crystal structures of mannose trimming enzyme drosophila Golgi alpha-mannosidase II (dGMII) complexed with the inhibitors mannostatin A (1) and an N-benzyl analogue (2) have been determined. Molecular dynamics simulations and NMR studies have shown that the five-membered ring of mannostatin A is rather flexible occupying pseudorotational itineraries between 2T3 and 5E, and 2T3 and 4E. In the bound state, mannostatin A adopts a 2T1 twist envelope conformation, which is not significantly populated in solution. Possible conformations of the mannosyl oxacarbenium ion and an enzyme-linked intermediate have been compared to the conformation of mannostatin A in the cocrystal structure with dGMII. It has been found that mannostatin A best mimics the covalent linked mannosyl intermediate, which adopts a 1S5 skew boat conformation. The thiomethyl group, which is critical for high affinity, superimposes with the C-6 hydroxyl of the covalent linked intermediate. This functionality is able to make a number of additional polar and nonpolar interactions increasing the affinity for dGMII. Furthermore, the X-ray structures show that the environment surrounding the thiomethyl group of 1 is remarkably similar to the arrangements around the methionine residues in the protein. Collectively, our studies contradict the long held view that potent inhibitors of glycosidases must mimic an oxacarbenium ion like transition state.
Organizational Affiliation:
Complex Carbohydrate Research Center, University of Georgia, 315 Riverbend Road, Athens, Georgia 30602, USA.