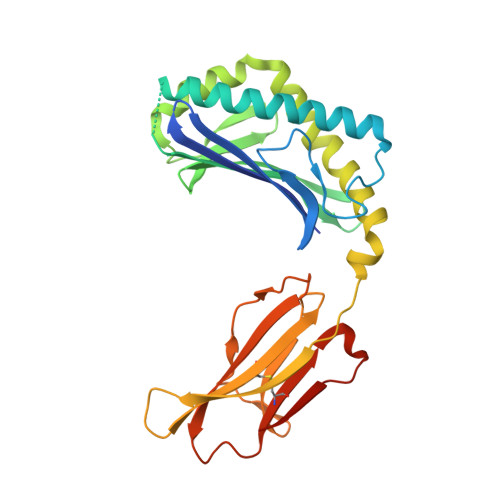
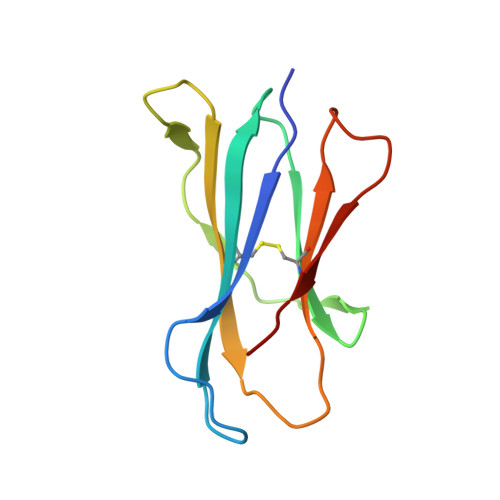
Design of natural killer T cell activators: structure and function of a microbial glycosphingolipid bound to mouse CD1d.
Wu, D., Zajonc, D.M., Fujio, M., Sullivan, B.A., Kinjo, Y., Kronenberg, M., Wilson, I.A., Wong, C.H.(2006) Proc Natl Acad Sci U S A 103: 3972-3977
- PubMed: 16537470
- DOI: https://doi.org/10.1073/pnas.0600285103
- Primary Citation of Related Structures:
2FIK - PubMed Abstract:
Natural killer T (NKT) cells provide an innate-type immune response upon T cell receptor interaction with CD1d-presented antigens. We demonstrate through equilibrium tetramer binding and antigen presentation assays with Valpha14i-positive NKT cell hybridomas that the Sphingomonas glycolipid alpha-galacturonosyl ceramide (GalA-GSL) is a NKT cell agonist that is significantly weaker than alpha-galactosylceramide (alpha-GalCer), the most potent known NKT agonist. For GalA-GSL, a shorter fatty acyl chain, an absence of the 4-OH on the sphingosine tail and a 6'-COOH group on the galactose moiety account for its observed antigenic potency. We further determined the crystal structure of mCD1d in complex with GalA-GSL at 1.8-A resolution. The overall binding mode of GalA-GSL to mCD1d is similar to that of the short-chain alpha-GalCer ligand PBS-25, but its sphinganine chain is more deeply inserted into the F' pocket due to alternate hydrogen-bonding interactions between the sphinganine 3-OH with Asp-80. Subsequently, a slight lateral shift (>1 A) of the galacturonosyl head group is noted at the CD1 surface compared with the galactose of alpha-GalCer. Because the relatively short C(14) fatty acid of GalA-GSL does not fully occupy the A' pocket, a spacer lipid is found that stabilizes this pocket. The lipid spacer was identified by GC/MS as a mixture of saturated and monounsaturated palmitic acid (C(16)). Comparison of available crystal structures of alpha-anomeric glycosphingolipids now sheds light on the structural basis of their differential antigenic potency and has led to the design and synthesis of NKT cell agonists with enhanced cell-based stimulatory activities compared with alpha-GalCer.
Organizational Affiliation:
Department of Chemistry and Molecular Biology and The Skaggs Institute for Chemical Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.