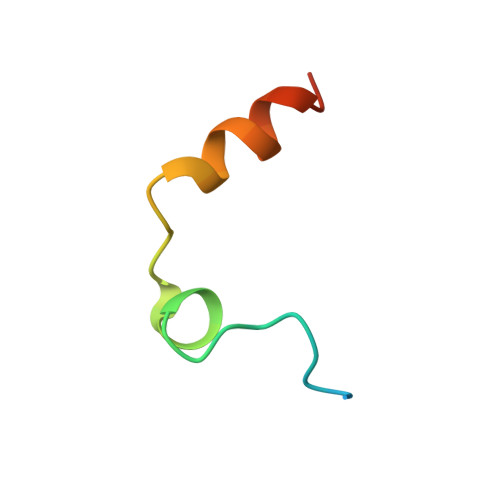
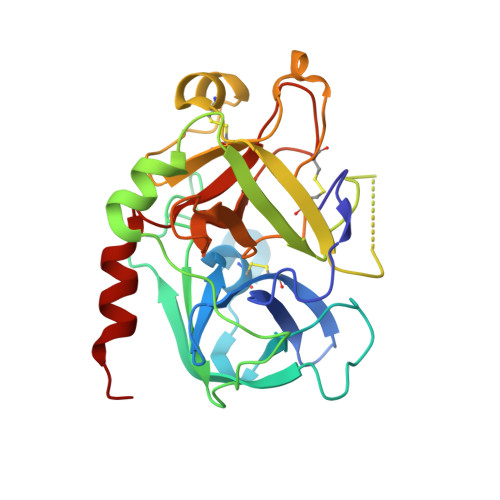
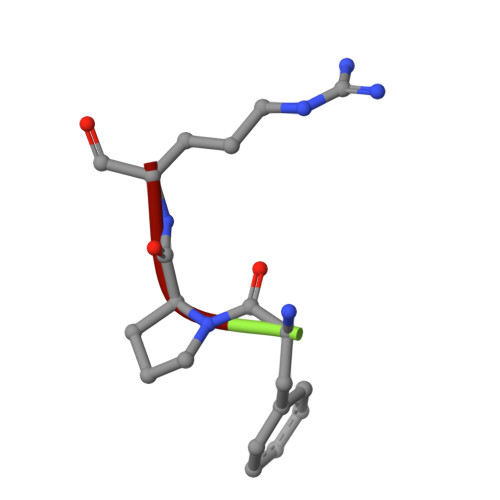
Structural and thermodynamic analysis of thrombin:suramin interaction in solution and crystal phases.
Lima, L.M., Becker, C.F., Giesel, G.M., Marques, A.F., Cargnelutti, M.T., de Oliveira Neto, M., Queiroz Monteiro, R., Verli, H., Polikarpov, I.(2009) Biochim Biophys Acta 1794: 873-881
- PubMed: 19332154
- DOI: https://doi.org/10.1016/j.bbapap.2009.03.011
- Primary Citation of Related Structures:
2H9T, 3BF6 - PubMed Abstract:
Suramin is a hexasulfonated naphthylurea which has been recently characterized as a non-competitive inhibitor of human alpha-thrombin activity over fibrinogen, although its binding site and mode of interaction with the enzyme remain elusive. Here, we determined two X-ray structure of the thrombin:suramin complex, refined at 2.4 A resolution. While a single thrombin:suramin complex was found in the asymmetric unit cell of the crystal, some of the crystallographic contacts with symmetrically related molecules are mediated by both the enzyme and the ligand. Molecular dynamics simulations with the 1:1 complex demonstrate a large rearrangement of suramin in the complex, but with the protein scaffold and the more extensive protein-ligand regions keep unchanged. Small-angle X-ray scattering measurements at high micromolar concentration demonstrate a suramin-induced dimerization of the enzyme. These data indicating a dissimilar binding mode in the monomeric and oligomeric states, with a monomeric, 1:1 complex to be more likely to exist at the thrombin physiological, nanomolar concentration range. Collectively, close understanding on the structural basis for interaction is given which might establish a basis for design of suramin analogues targeting thrombin.
Organizational Affiliation:
Faculdade de Farmácia, Universidade Federal do Rio de Janeiro, 21941-590, Rio de Janeiro, RJ, Brazil. [email protected]