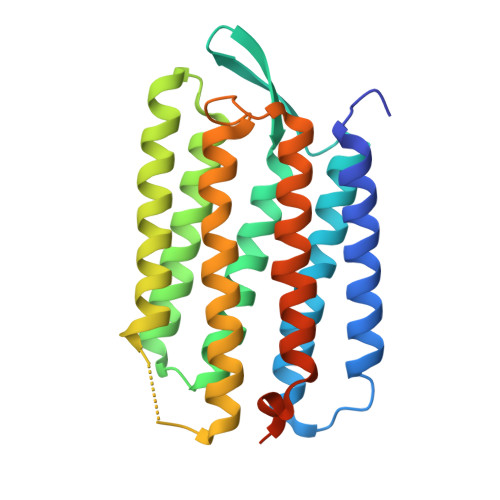
Structural changes in the L photointermediate of bacteriorhodopsin.
Lanyi, J.K., Schobert, B.(2007) J Mol Biol 365: 1379-1392
- PubMed: 17141271
- DOI: https://doi.org/10.1016/j.jmb.2006.11.016
- Primary Citation of Related Structures:
2NTU, 2NTW - PubMed Abstract:
The L to M reaction of the bacteriorhodopsin photocycle includes the crucial proton transfer from the retinal Schiff base to Asp85. In spite of the importance of the L state in deciding central issues of the transport mechanism in this pump, the serious disagreements among the three published crystallographic structures of L have remained unresolved. Here, we report on the X-ray diffraction structure of the L state, to 1.53-1.73 A resolutions, from replicate data sets collected from six independent crystals. Unlike earlier studies, the partial occupancy refinement uses diffraction intensities from the same crystals before and after the illumination to produce the trapped L state. The high reproducibility of inter-atomic distances, and bond angles and torsions of the retinal, lends credibility to the structural model. The photoisomerized 13-cis retinal in L is twisted at the C(13)=C(14) and C(15)=NZ double-bonds, and the Schiff base does not lose its connection to Wat402 and, therefore, to the proton acceptor Asp85. The protonation of Asp85 by the Schiff base in the L-->M reaction is likely to occur, therefore, via Wat402. It is evident from the structure of the L state that various conformational changes involving hydrogen-bonding residues and bound water molecules begin to propagate from the retinal to the protein at this stage already, and in both extracellular and cytoplasmic directions. Their rationales in the transport can be deduced from the way their amplitudes increase in the intermediates that follow L in the reaction cycle, and from the proton transfer reactions with which they are associated.
Organizational Affiliation:
Department of Physiology & Biophysics, University of California, Irvine, CA 92697, USA. [email protected]