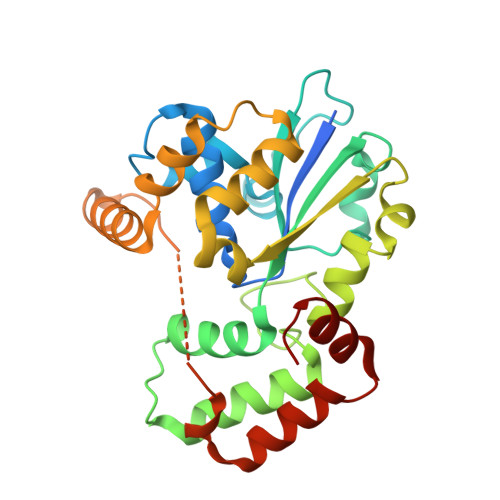
Crystal structure of StaL, a glycopeptide antibiotic sulfotransferase from Streptomyces toyocaensis.
Shi, R., Lamb, S.S., Bhat, S., Sulea, T., Wright, G.D., Matte, A., Cygler, M.(2007) J Biol Chem 282: 13073-13086
- PubMed: 17329243
- DOI: https://doi.org/10.1074/jbc.M611912200
- Primary Citation of Related Structures:
2OV8, 2OVB, 2OVF - PubMed Abstract:
Over the past decade, antimicrobial resistance has emerged as a major public health crisis. Glycopeptide antibiotics such as vancomycin and teicoplanin are clinically important for the treatment of Gram-positive bacterial infections. StaL is a 3'-phosphoadenosine 5'-phosphosulfate-dependent sulfotransferase capable of sulfating the cross-linked heptapeptide substrate both in vivo and in vitro, yielding the product A47934, a unique teicoplanin-class glycopeptide antibiotic. The sulfonation reaction catalyzed by StaL constitutes the final step in A47934 biosynthesis. Here we report the crystal structure of StaL and its complex with the cofactor product 3'-phosphoadenosine 5'-phosphate. This is only the second prokaryotic sulfotransferase to be structurally characterized. StaL belongs to the large sulfotransferase family and shows higher similarity to cytosolic sulfotransferases (ST) than to the bacterial ST (Stf0). StaL has a novel dimerization motif, different from any other STs that have been structurally characterized. We have also applied molecular modeling to investigate the binding mode of the unique substrate, desulfo-A47934. Based on the structural analysis and modeling results, a series of residues was mutated and kinetically characterized. In addition to the conserved residues (Lys(12), His(67), and Ser(98)), molecular modeling, fluorescence quenching experiments, and mutagenesis studies identified several other residues essential for substrate binding and/or activity, including Trp(34), His(43), Phe(77), Trp(132), and Glu(205).
Organizational Affiliation:
Department of Biochemistry, McGill University, Montréal, Québec H3G 1Y6.