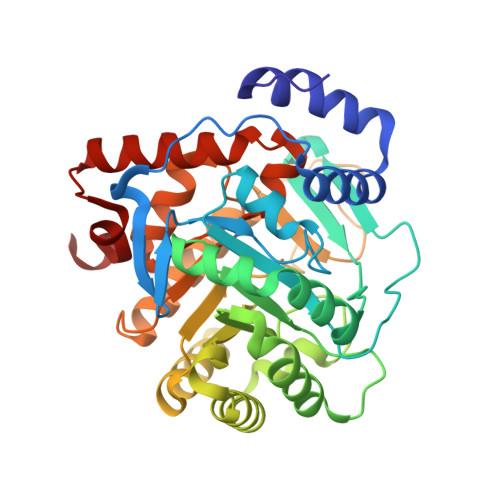
The structures of human dihydroorotate dehydrogenase with and without inhibitor reveal conformational flexibility in the inhibitor and substrate binding sites
Walse, B., Dufe, V.T., Svensson, B., Fritzson, I., Dahlberg, L., Khairoullina, A., Wellmar, U., Al-Karadaghi, S.(2008) Biochemistry 47: 8929-8936
- PubMed: 18672895
- DOI: https://doi.org/10.1021/bi8003318
- Primary Citation of Related Structures:
2PRH, 2PRL, 2PRM - PubMed Abstract:
Inhibitors of dihydroorotate dehydrogenase (DHODH) have been suggested for the treatment of rheumatoid arthritis, psoriasis, autoimmune diseases, Plasmodium, and bacterial and fungal infections. Here we present the structures of N-terminally truncated (residues Met30-Arg396) DHODH in complex with two inhibitors: a brequinar analogue (6) and a novel inhibitor (a fenamic acid derivative) (7), as well as the first structure of the enzyme to be characterized without any bound inhibitor. It is shown that 7 uses the "standard" brequinar binding mode and, in addition, interacts with Tyr356, a residue conserved in most class 2 DHODH proteins. Compared to the inhibitor-free structure, some of the amino acid side chains in the tunnel in which brequinar binds and which was suggested to be the binding site of ubiquinone undergo changes in conformation upon inhibitor binding. Using our data, the loop regions of residues Leu68-Arg72 and Asn212-Leu224, which were disordered in previously studied human DHODH structures, could be built into the electron density. The first of these loops, which is located at the entrance to the inhibitor-binding pocket, shows different conformations in the three structures, suggesting that it may interfere with inhibitor/cofactor binding. The second loop has been suggested to control the access of dihydroorotate to the active site of the enzyme and may be an important player in the enzymatic reaction. These observations provide new insights into the dynamic features of the DHODH reaction and suggest new approaches to the design of inhibitors against DHODH.
Organizational Affiliation:
SARomics AB, P.O. Box 724, SE-220 07 Lund, Sweden. [email protected]