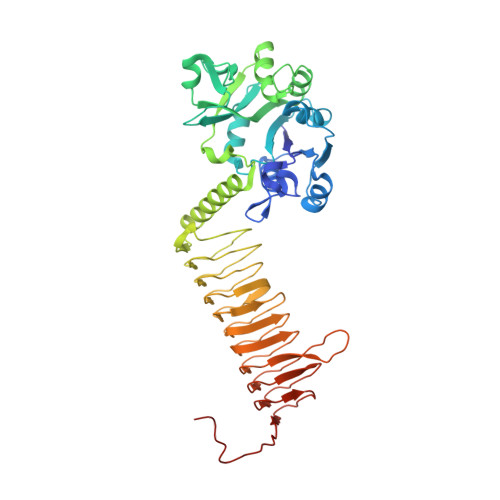
Characterization of Substrate Binding and Catalysis in the Potential Antibacterial Target N-Acetylglucosamine-1-Phosphate Uridyltransferase (Glmu).
Mochalkin, I., Lightle, S., Zhu, Y., Ohren, J.F., Spessard, C., Chirgadze, N.Y., Banotai, C., Melnick, M., Mcdowell, L.(2007) Protein Sci 16: 2657
- PubMed: 18029420
- DOI: https://doi.org/10.1110/ps.073135107
- Primary Citation of Related Structures:
2V0H, 2V0I, 2V0J, 2V0K, 2V0L - PubMed Abstract:
N-Acetylglucosamine-1-phosphate uridyltransferase (GlmU) catalyzes the first step in peptidoglycan biosynthesis in both Gram-positive and Gram-negative bacteria. The products of the GlmU reaction are essential for bacterial survival, making this enzyme an attractive target for antibiotic drug discovery. A series of Haemophilus influenzae GlmU (hiGlmU) structures were determined by X-ray crystallography in order to provide structural and functional insights into GlmU activity and inhibition. The information derived from these structures was combined with biochemical characterization of the K25A, Q76A, D105A, Y103A, V223A, and E224A hiGlmU mutants in order to map these active-site residues to catalytic activity of the enzyme and refine the mechanistic model of the GlmU uridyltransferase reaction. These studies suggest that GlmU activity follows a sequential substrate-binding order that begins with UTP binding noncovalently to the GlmU enzyme. The uridyltransferase active site then remains in an open apo-like conformation until N-acetylglucosamine-1-phosphate (GlcNAc-1-P) binds and induces a conformational change at the GlcNAc-binding subsite. Following the binding of GlcNAc-1-P to the UTP-charged uridyltransferase active site, the non-esterified oxygen of GlcNAc-1-P performs a nucleophilic attack on the alpha-phosphate group of UTP. The new data strongly suggest that the mechanism of phosphotransfer in the uridyltransferase reaction in GlmU is primarily through an associative mechanism with a pentavalent phosphate intermediate and an inversion of stereochemistry. Finally, the structural and biochemical characterization of the uridyltransferase active site and catalytic mechanism described herein provides a basis for the structure-guided design of novel antibacterial agents targeting GlmU activity.
Organizational Affiliation:
Pfizer Global Research and Development, Michigan Laboratories, Ann Arbor, Michigan 48105, USA. [email protected]