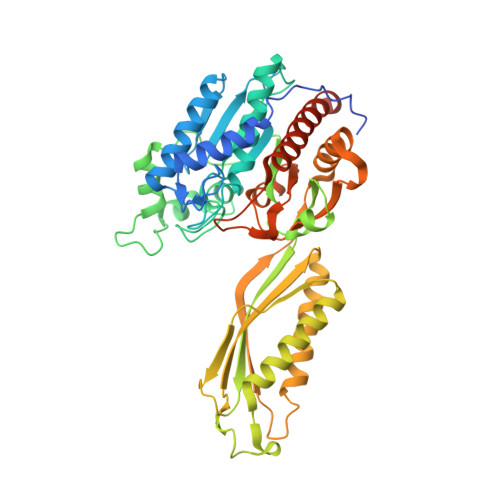
Crystal Structures of Yeast -Alanine Synthase Complexes Reveal the Mode of Substrate Binding and Large Scale Domain Closure Movements.
Lundgren, S., Andersen, B., Piskur, J., Dobritzsch, D.(2007) J Biol Chem 282: 36037
- PubMed: 17916556
- DOI: https://doi.org/10.1074/jbc.M705517200
- Primary Citation of Related Structures:
2V8D, 2V8G, 2V8H, 2V8V - PubMed Abstract:
Beta-alanine synthase is the final enzyme of the reductive pyrimidine catabolic pathway, which is responsible for the breakdown of uracil and thymine in higher organisms. The fold of the homodimeric enzyme from the yeast Saccharomyces kluyveri identifies it as a member of the AcyI/M20 family of metallopeptidases. Its subunit consists of a catalytic domain harboring a di-zinc center and a smaller dimerization domain. The present site-directed mutagenesis studies identify Glu(159) and Arg(322) as crucial for catalysis and His(262) and His(397) as functionally important but not essential. We determined the crystal structures of wild-type beta-alanine synthase in complex with the reaction product beta-alanine, and of the mutant E159A with the substrate N-carbamyl-beta-alanine, revealing the closed state of a dimeric AcyI/M20 metallopeptidase-like enzyme. Subunit closure is achieved by a approximately 30 degrees rigid body domain rotation, which completes the active site by integration of substrate binding residues that belong to the dimerization domain of the same or the partner subunit. Substrate binding is achieved via a salt bridge, a number of hydrogen bonds, and coordination to one of the zinc ions of the di-metal center.
Organizational Affiliation:
Department of Medical Biochemistry and Biophysics, Karolinska Institutet, SE-17177 Stockholm, Sweden.