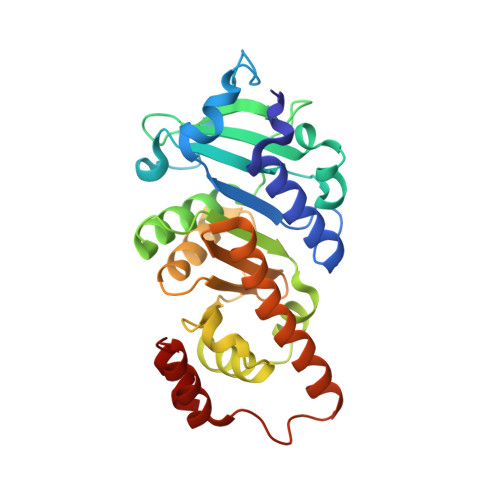
Designed Protein-Protein Association.
Grueninger, D., Treiber, N., Ziegler, M.O.P., Koetter, J.W.A., Schulze, M.-S., Schulz, G.E.(2008) Science 319: 206
- PubMed: 18187656
- DOI: https://doi.org/10.1126/science.1150421
- Primary Citation of Related Structures:
2UYU, 2UYV, 2V7G, 2V9E, 2V9F, 2V9G, 2V9I, 2V9L, 2V9M, 2V9N, 2V9O, 2V9U - PubMed Abstract:
The analysis of natural contact interfaces between protein subunits and between proteins has disclosed some general rules governing their association. We have applied these rules to produce a number of novel assemblies, demonstrating that a given protein can be engineered to form contacts at various points of its surface. Symmetry plays an important role because it defines the multiplicity of a designed contact and therefore the number of required mutations. Some of the proteins needed only a single side-chain alteration in order to associate to a higher-order complex. The mobility of the buried side chains has to be taken into account. Four assemblies have been structurally elucidated. Comparisons between the designed contacts and the results will provide useful guidelines for the development of future architectures.
Organizational Affiliation:
Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, Albertstrasse 21, 79104 Freiburg im Breisgau, Germany.