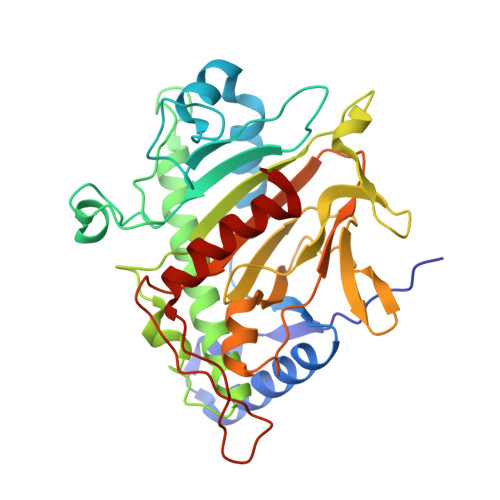
The Crystal Structure of Isopenicillin N Synthase with Delta((L)-Alpha-Aminoadipoyl)-(L)-Cysteinyl-(D)-Methionine Reveals Thioether Coordination to Iron.
Clifton, I.J., Ge, W., Adlington, R.M., Baldwin, J.E., Rutledge, P.J.(2011) Arch Biochem Biophys 516: 103
- PubMed: 22001738
- DOI: https://doi.org/10.1016/j.abb.2011.09.014
- Primary Citation of Related Structures:
2Y60 - PubMed Abstract:
Isopenicillin N synthase (IPNS) catalyses cyclization of δ-(l-α-aminoadipoyl)-l-cysteinyl-d-valine (ACV) to isopenicillin N (IPN), the central step in penicillin biosynthesis. Previous studies have shown that IPNS turns over a wide range of substrate analogues in which the valine residue of its natural substrate is replaced with other amino acids. IPNS accepts and oxidizes numerous substrates that bear hydrocarbon sidechains in this position, however the enzyme is less tolerant of analogues presenting polar functionality in place of the valinyl isopropyl group. We report a new ACV analogue δ-(l-α-aminoadipoyl)-l-cysteinyl-d-methionine (ACM), which incorporates a thioether in place of the valinyl sidechain. ACM has been synthesized using solution phase methods and crystallized with IPNS. A crystal structure has been elucidated for the IPNS:Fe(II):ACM complex at 1.40Å resolution. This structure reveals that ACM binds in the IPNS active site such that the sulfur atom of the methionine thioether binds to iron in the oxygen binding site at a distance of 2.57Å. The sulfur of the cysteinyl thiolate sits 2.36Å from the metal.
Organizational Affiliation:
Chemistry Research Laboratory, University of Oxford, UK.