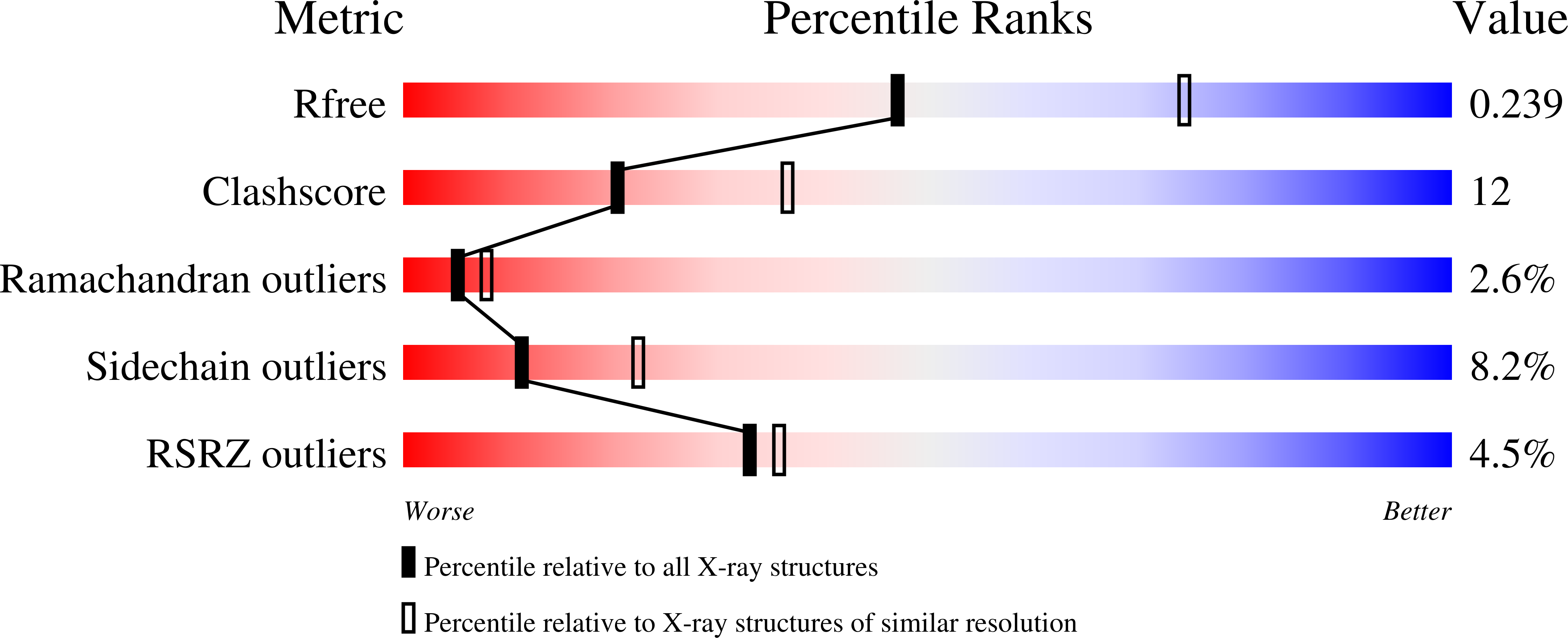
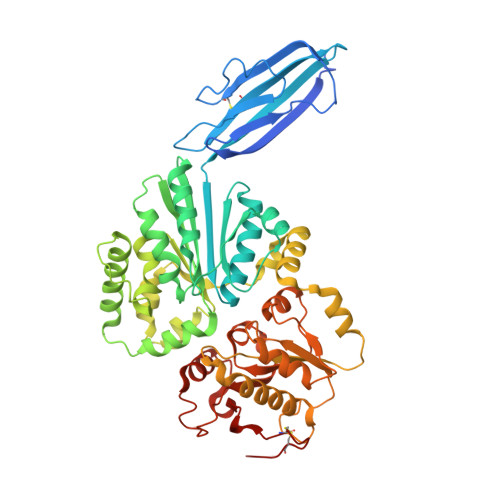
Crystal structure of Vibrionaceae Photobacterium sp. JT-ISH-224 alpha2,6-sialyltransferase in a ternary complex with donor product CMP and acceptor substrate lactose: catalytic mechanism and substrate recognition
Kakuta, Y., Okino, N., Kajiwara, H., Ichikawa, M., Takakura, Y., Ito, M., Yamamoto, T.(2008) Glycobiology 18: 66-73
- PubMed: 17962295
- DOI: https://doi.org/10.1093/glycob/cwm119
- Primary Citation of Related Structures:
2Z4T - PubMed Abstract:
Sialyltransferases are a family of glycosyltransferases that catalyze the transfer of N-acetylneuraminic acid residues from cytidine monophosphate N-acetylneuraminic acid (CMP-NeuAc) as a donor substrate to the carbohydrate groups of glycoproteins and glycolipids as acceptor substrates. We determined the crystal structure of Delta16psp26ST, the N-terminal truncated form of alpha2,6-sialyltransferase from Vibrionaceae Photobacterium sp. JT-ISH-224, complexed with a donor product CMP and an acceptor substrate lactose. Delta16psp26ST has three structural domains. Domain 1 belongs to the immunoglobulin-like beta-sandwich fold, and domains 2 and 3 form the glycosyltransferase-B structure. The CMP and lactose were bound in the deep cleft between domains 2 and 3. In the structure, only Asp232 was within hydrogen-binding distance of the acceptor O6 carbon of the galactose residue in lactose, and His405 was within hydrogen-binding distance of the phosphate oxygen of CMP. Mutation of these residues greatly decreased the activity of the enzyme. These structural and mutational results indicated that Asp232 might act as a catalytic base for deprotonation of the acceptor substrate, and His405 might act as a catalytic acid for protonation of the donor substrate. These findings are consistent with an in-line-displacement reaction mechanism in which Delta16psp26ST catalyzes the inverting transfer reaction. Unlike the case with multifunctional sialyltransferase (Delta24PmST1) complexed with CMP and lactose, the crystal structure of which was recently reported, the alpha2,6 reaction specificity of Delta16psp26ST is likely to be determined by His123.
Organizational Affiliation:
Department of Bioscience and Biotechnology, Graduate School of Bioresource and Bioenvironmental Sciences, Kyushu University, 6-10-1 Hakozaki, Higashi-ku, Fukuoka 812-8581, Japan. [email protected]