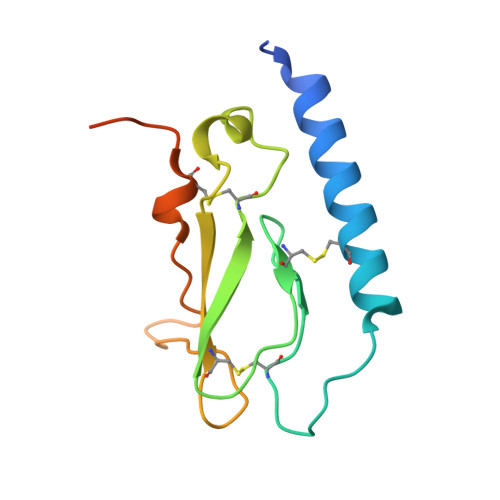
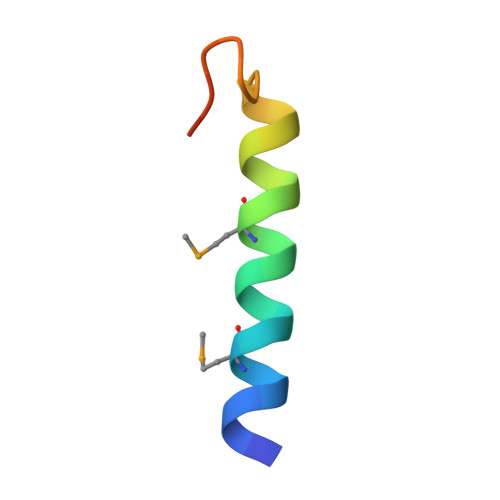
Crystal Structure of the Ligand-bound Glucagon-like Peptide-1 Receptor Extracellular Domain
Runge, S., Thogersen, H., Madsen, K., Lau, J., Rudolph, R.(2008) J Biol Chem 283: 11340-11347
- PubMed: 18287102
- DOI: https://doi.org/10.1074/jbc.M708740200
- Primary Citation of Related Structures:
3C59, 3C5T - PubMed Abstract:
The glucagon-like peptide-1 receptor (GLP-1R) belongs to Family B1 of the seven-transmembrane G protein-coupled receptors, and its natural agonist ligand is the peptide hormone glucagon-like peptide-1 (GLP-1). GLP-1 is involved in glucose homeostasis, and activation of GLP-1R in the plasma membrane of pancreatic beta-cells potentiates glucose-dependent insulin secretion. The N-terminal extracellular domain (nGLP-1R) is an important ligand binding domain that binds GLP-1 and the homologous peptide Exendin-4 with differential affinity. Exendin-4 has a C-terminal extension of nine amino acid residues known as the "Trp cage", which is absent in GLP-1. The Trp cage was believed to interact with nGLP-1R and thereby explain the superior affinity of Exendin-4. However, the molecular details that govern ligand binding and specificity of nGLP-1R remain undefined. Here we report the crystal structure of human nGLP-1R in complex with the antagonist Exendin-4(9-39) solved by the multiwavelength anomalous dispersion method to 2.2A resolution. The structure reveals that Exendin-4(9-39) is an amphipathic alpha-helix forming both hydrophobic and hydrophilic interactions with nGLP-1R. The Trp cage of Exendin-4 is not involved in binding to nGLP-1R. The hydrophobic binding site of nGLP-1R is defined by discontinuous segments including primarily a well defined alpha-helix in the N terminus of nGLP-1R and a loop between two antiparallel beta-strands. The structure provides for the first time detailed molecular insight into ligand binding of the human GLP-1 receptor, an established target for treatment of type 2 diabetes.
Organizational Affiliation:
Department of Structure and Biophysical Chemistry, Novo Nordisk, 2760 Måløv, Denmark. [email protected]