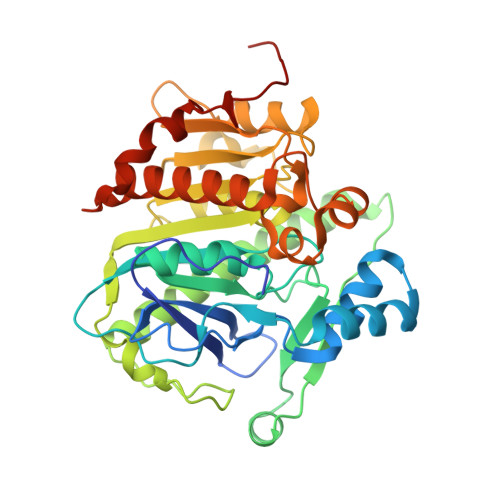
Crystal Structure of Human Plasma Platelet-activating Factor Acetylhydrolase: STRUCTURAL IMPLICATION TO LIPOPROTEIN BINDING AND CATALYSIS.
Samanta, U., Bahnson, B.J.(2008) J Biol Chem 283: 31617-31624
- PubMed: 18784071
- DOI: https://doi.org/10.1074/jbc.M804750200
- Primary Citation of Related Structures:
3D59, 3D5E - PubMed Abstract:
Human plasma platelet-activating factor (PAF) acetylhydrolase functions by reducing PAF levels as a general anti-inflammatory scavenger and is linked to anaphylactic shock, asthma, and allergic reactions. The enzyme has also been implicated in hydrolytic activities of other pro-inflammatory agents, such as sn-2 oxidatively fragmented phospholipids. This plasma enzyme is tightly bound to low and high density lipoprotein particles and is also referred to as lipoprotein-associated phospholipase A2. The crystal structure of this enzyme has been solved from x-ray diffraction data collected to a resolution of 1.5 angstroms. It has a classic lipase alpha/beta-hydrolase fold, and it contains a catalytic triad of Ser273, His351, and Asp296. Two clusters of hydrophobic residues define the probable interface-binding region, and a prediction is given of how the enzyme is bound to lipoproteins. Additionally, an acidic patch of 10 carboxylate residues and a neighboring basic patch of three residues are suggested to play a role in high density lipoprotein/low density lipoprotein partitioning. A crystal structure is also presented of PAF acetylhydrolase reacted with the organophosphate compound paraoxon via its active site Ser273. The resulting diethyl phosphoryl complex was used to model the tetrahedral intermediate of the substrate PAF to the active site. The model of interface binding begins to explain the known specificity of lipoprotein-bound substrates and how the active site can be both close to the hydrophobic-hydrophilic interface and at the same time be accessible to the aqueous phase.
Organizational Affiliation:
Department of Chemistry & Biochemistry, University of Delaware, Newark, Delaware 19716, USA.