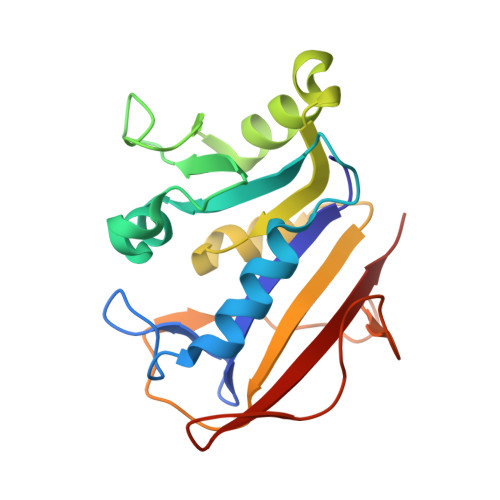
Structural analysis of a holoenzyme complex of mouse dihydrofolate reductase with NADPH and a ternary complex with the potent and selective inhibitor 2,4-diamino-6-(2'-hydroxydibenz[b,f]azepin-5-yl)methylpteridine.
Cody, V., Pace, J., Rosowsky, A.(2008) Acta Crystallogr D Biol Crystallogr 64: 977-984
- PubMed: 18703847
- DOI: https://doi.org/10.1107/S0907444908022348
- Primary Citation of Related Structures:
3D80, 3D84 - PubMed Abstract:
It has been shown that 2,4-diamino-6-arylmethylpteridines and 2,4-diamino-5-arylmethylpyrimidines containing an O-carboxylalkyloxy group in the aryl moiety are potent and selective inhibitors of the dihydrofolate reductase (DHFR) from opportunistic pathogens such as Pneumocystis carinii, the causative agent of Pneumocystis pneumonia in HIV/AIDS patients. In order to understand the structure-activity profile observed for a series of substituted dibenz[b,f]azepine antifolates, the crystal structures of mouse DHFR (mDHFR; a mammalian homologue) holo and ternary complexes with NADPH and the inhibitor 2,4-diamino-6-(2'-hydroxydibenz[b,f]azepin-5-yl)methylpteridine were determined to 1.9 and 1.4 A resolution, respectively. Structural data for the ternary complex with the potent O-(3-carboxypropyl) inhibitor PT684 revealed no electron density for the O-carboxylalkyloxy side chain. The side chain was either cleaved or completely disordered. The electron density fitted the less potent hydroxyl compound PT684a. Additionally, cocrystallization of mDHFR with NADPH and the less potent 2'-(4-carboxybenzyl) inhibitor PT682 showed no electron density for the inhibitor and resulted in the first report of a holoenzyme complex despite several attempts at crystallization of a ternary complex. Modeling data of PT682 in the active site of mDHFR and P. carinii DHFR (pcDHFR) indicate that binding would require ligand-induced conformational changes to the enzyme for the inhibitor to fit into the active site or that the inhibitor side chain would have to adopt an alternative binding mode to that observed for other carboxyalkyloxy inhibitors. These data also show that the mDHFR complexes have a decreased active-site volume as reflected in the relative shift of helix C (residues 59-64) by 0.6 A compared with pcDHFR ternary complexes. These data are consistent with the greater inhibitory potency against pcDHFR.
Organizational Affiliation:
Structural Biology Department, Hauptman-Woodward Medical Research Institute, 700 Ellicott Street, Buffalo, NY 14203, USA. [email protected]