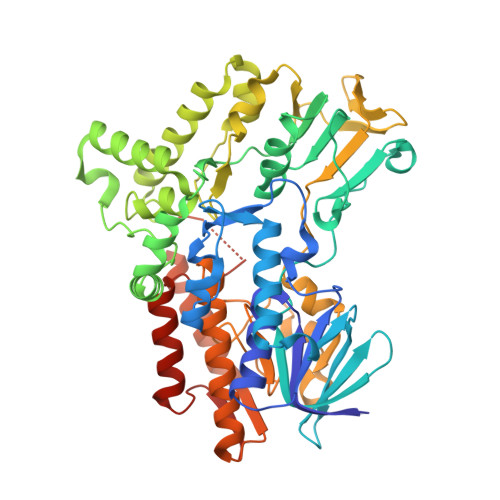
Crystal structures of cyclohexanone monooxygenase reveal complex domain movements and a sliding cofactor
Mirza, I.A., Yachnin, B.J., Wang, S., Grosse, S., Bergeron, H., Imura, A., Iwaki, H., Hasegawa, Y., Lau, P.C., Berghuis, A.M.(2009) J Am Chem Soc 131: 8848-8854
- PubMed: 19385644
- DOI: https://doi.org/10.1021/ja9010578
- Primary Citation of Related Structures:
3GWD, 3GWF - PubMed Abstract:
Cyclohexanone monooxygenase (CHMO) is a flavoprotein that carries out the archetypical Baeyer-Villiger oxidation of a variety of cyclic ketones into lactones. Using NADPH and O(2) as cosubstrates, the enzyme inserts one atom of oxygen into the substrate in a complex catalytic mechanism that involves the formation of a flavin-peroxide and Criegee intermediate. We present here the atomic structures of CHMO from an environmental Rhodococcus strain bound with FAD and NADP(+) in two distinct states, to resolutions of 2.3 and 2.2 A. The two conformations reveal domain shifts around multiple linkers and loop movements, involving conserved arginine 329 and tryptophan 492, which effect a translation of the nicotinamide resulting in a sliding cofactor. Consequently, the cofactor is ideally situated and subsequently repositioned during the catalytic cycle to first reduce the flavin and later stabilize formation of the Criegee intermediate. Concurrent movements of a loop adjacent to the active site demonstrate how this protein can effect large changes in the size and shape of the substrate binding pocket to accommodate a diverse range of substrates. Finally, the previously identified BVMO signature sequence is highlighted for its role in coordinating domain movements. Taken together, these structures provide mechanistic insights into CHMO-catalyzed Baeyer-Villiger oxidation.
Organizational Affiliation:
Department of Biochemistry, McGill University, 3649 Prom Sir William Osler, Bellini Pavilion, Room 466, Montreal, QC, Canada H3G 0B1.