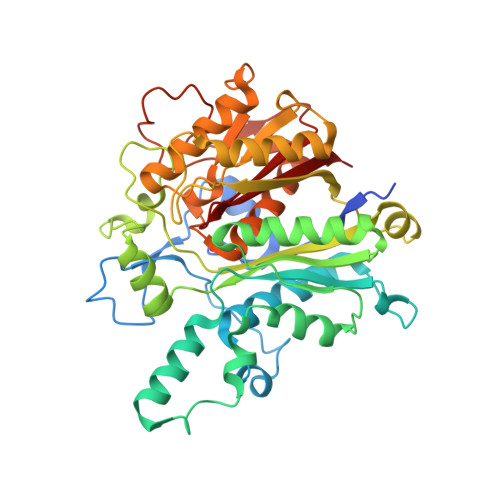
Isolation, enzyme-bound structure and antibacterial activity of platencin A1 from Streptomyces platensis.
Singh, S.B., Ondeyka, J.G., Herath, K.B., Zhang, C., Jayasuriya, H., Zink, D.L., Parthasarathy, G., Becker, J.W., Wang, J., Soisson, S.M.(2009) Bioorg Med Chem Lett 19: 4756-4759
- PubMed: 19581087
- DOI: https://doi.org/10.1016/j.bmcl.2009.06.061
- Primary Citation of Related Structures:
3HNZ, 3HO2, 3HO9, 3I8P - PubMed Abstract:
Natural products continue to serve as one of the best sources for discovery of antibacterial agents as exemplified by the recent discoveries of platensimycin and platencin. Chemical modifications as well as discovery of congeners are the main sources for gaining knowledge of structure-activity relationship of natural products. Screening for congeners in the extracts of the fermentation broths of Streptomyces platensis led to the isolation of platencin A(1), a hydroxy congener of platencin. The hydroxylation of the tricyclic enone moiety negatively affected the antibacterial activity and appears to be consistent with the hydrophobic binding pocket of the FabF. Isolation, structure, enzyme-bound structure and activity of platencin A(1) and two other congeners have been described.
Organizational Affiliation:
Merck Research Laboratories, Rahway, NJ 07065, USA. [email protected]